

You can:
| Name | C-X-C chemokine receptor type 5 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CXCR5 |
| Synonym | BLR-1 NLR neurolymphatic receptor Monocyte-derived receptor 15 MDR15 [ Show all ] |
| Disease | Systemic lupus erythematosus |
| Length | 372 |
| Amino acid sequence | MNYPLTLEMDLENLEDLFWELDRLDNYNDTSLVENHLCPATEGPLMASFKAVFVPVAYSLIFLLGVIGNVLVLVILERHRQTRSSTETFLFHLAVADLLLVFILPFAVAEGSVGWVLGTFLCKTVIALHKVNFYCSSLLLACIAVDRYLAIVHAVHAYRHRRLLSIHITCGTIWLVGFLLALPEILFAKVSQGHHNNSLPRCTFSQENQAETHAWFTSRFLYHVAGFLLPMLVMGWCYVGVVHRLRQAQRRPQRQKAVRVAILVTSIFFLCWSPYHIVIFLDTLARLKAVDNTCKLNGSLPVAITMCEFLGLAHCCLNPMLYTFAGVKFRSDLSRLLTKLGCTGPASLCQLFPSWRRSSLSESENATSLTTF |
| UniProt | P32302 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P32302 |
| 3D structure model | This predicted structure model is from GPCR-EXP P32302. |
| BioLiP | N/A |
| Therapeutic Target Database | T09528 |
| ChEMBL | CHEMBL1075315 |
| IUPHAR | N/A |
| DrugBank | N/A |
| Name | CHEMBL2179458 |
|---|---|
| Molecular formula | C174H174F108N30O12P18 |
| IUPAC name | 1-[3-[4-[1-[[3-[[4-[1-[[3,5-bis[[4-[1-[[3,5-bis[[4-[1-[3-(5-methyl-2,4-dioxopyrimidin-1-yl)propyl]pyridin-1-ium-4-yl]pyridin-1-ium-1-yl]methyl]phenyl]methyl]pyridin-1-ium-4-yl]pyridin-1-ium-1-yl]methyl]phenyl]methyl]pyridin-1-ium-4-yl]pyridin-1-ium-1-yl]methyl]-5-[[4-[1-[3-(5-methyl-2,4-dioxopyrimidin-1-yl)propyl]pyridin-1-ium-4-yl]pyridin-1-ium-1-yl]methyl]phenyl]methyl]pyridin-1-ium-4-yl]pyridin-1-ium-1-yl]propyl]-5-methylpyrimidine-2,4-dione;octadecahexafluorophosphate |
| Molecular weight | 5486.86 |
| Hydrogen bond acceptor | 138 |
| Hydrogen bond donor | 6 |
| XlogP | None |
| Synonyms | N/A |
| Inchi Key | IZRYPUTVERTOHV-UHFFFAOYSA-T |
| Inchi ID | InChI=1S/C174H168N30O12.18F6P/c1-127-109-199(169(211)175-163(127)205)55-7-49-181-61-13-145(14-62-181)151-25-73-187(74-26-151)115-133-97-134(116-188-75-27-152(28-76-188)146-15-63-182(64-16-146)50-8-56-200-110-128(2)164(206)176-170(200)212)101-139(100-133)121-193-85-37-157(38-86-193)160-43-91-196(92-44-160)124-142-106-143(125-197-93-45-161(46-94-197)158-39-87-194(88-40-158)122-140-102-135(117-189-77-29-153(30-78-189)147-17-65-183(66-18-147)51-9-57-201-111-129(3)165(207)177-171(201)213)98-136(103-140)118-190-79-31-154(32-80-190)148-19-67-184(68-20-148)52-10-58-202-112-130(4)166(208)178-172(202)214)108-144(107-142)126-198-95-47-162(48-96-198)159-41-89-195(90-42-159)123-141-104-137(119-191-81-33-155(34-82-191)149-21-69-185(70-22-149)53-11-59-203-113-131(5)167(209)179-173(203)215)99-138(105-141)120-192-83-35-156(36-84-192)150-23-71-186(72-24-150)54-12-60-204-114-132(6)168(210)180-174(204)216;18*1-7(2,3,4,5)6/h13-48,61-114H,7-12,49-60,115-126H2,1-6H3;;;;;;;;;;;;;;;;;;/q+12;18*-1/p+6 |
| PubChem CID | 71457283 |
| ChEMBL | CHEMBL2179458 |
| IUPHAR | N/A |
| BindingDB | N/A |
| DrugBank | N/A |
Structure | 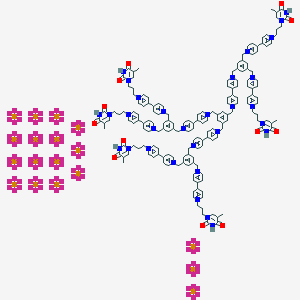 |
| Lipinski's druglikeness | This ligand has more than 5 hydrogen bond donor. This ligand has more than 10 hydrogen bond acceptor. This ligand is heavier than 500 daltons. Partition coefficient log P of this ligand is not available. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| IC50 | 10.25 ug.mL-1 | PMID23157587 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218