

You can:
| Name | Adenosine receptor A1 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Adora1 |
| Synonym | A1 receptor A1-AR A1R adenosine receptor A1 RDC7 |
| Disease | N/A for non-human GPCRs |
| Length | 326 |
| Amino acid sequence | MPPYISAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQTYFHTCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKTVVTQRRAAVAIAGCWILSLVVGLTPMFGWNNLSVVEQDWRANGSVGEPVIKCEFEKVISMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLNKKVSASSGDPQKYYGKELKIAKSLALILFLFALSWLPLHILNCITLFCPTCQKPSILIYIAIFLTHGNSAMNPIVYAFRIHKFRVTFLKIWNDHFRCQPKPPIDEDLPEEKAED |
| UniProt | P25099 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL318 |
| IUPHAR | 18 |
| DrugBank | N/A |
| Name | NECA |
|---|---|
| Molecular formula | C12H16N6O4 |
| IUPAC name | (2S,3S,4R,5R)-5-(6-aminopurin-9-yl)-N-ethyl-3,4-dihydroxyoxolane-2-carboxamide |
| Molecular weight | 308.298 |
| Hydrogen bond acceptor | 8 |
| Hydrogen bond donor | 4 |
| XlogP | -0.7 |
| Synonyms | Adenosine-5-(N-ethylcarboxamide) CS-8131 MFCD00069195 NCGC00025260-03 SCHEMBL16585523 [ Show all ] |
| Inchi Key | JADDQZYHOWSFJD-FLNNQWSLSA-N |
| Inchi ID | InChI=1S/C12H16N6O4/c1-2-14-11(21)8-6(19)7(20)12(22-8)18-4-17-5-9(13)15-3-16-10(5)18/h3-4,6-8,12,19-20H,2H2,1H3,(H,14,21)(H2,13,15,16)/t6-,7+,8-,12+/m0/s1 |
| PubChem CID | 448222 |
| ChEMBL | CHEMBL464859 |
| IUPHAR | 377, 425 |
| BindingDB | 21220 |
| DrugBank | N/A |
Structure | 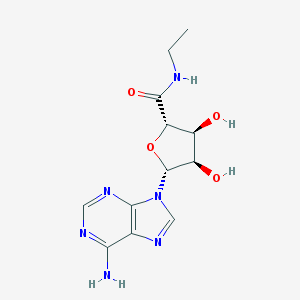 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | 54.8 nM | PMID8863798, PMID8201607 | ChEMBL |
| EC50 | 55.0 nM | PMID8863798, PMID8201607, PMID7658444 | BindingDB,ChEMBL |
| EC50 | 87.0 nM | PMID9871584 | BindingDB |
| EC50 | 87.2 nM | PMID9871584 | ChEMBL |
| EC50 | 54800.0 nM | PMID7739005 | BindingDB,ChEMBL |
| IC50 | 2.4 nM | PMID3351850 | BindingDB,ChEMBL |
| IC50 | 10.0 nM | PMID2374150 | BindingDB,ChEMBL |
| IC50 | 10.2 nM | PMID3018244 | BindingDB,ChEMBL |
| IC50 | 16.0 nM | PMID1875349 | BindingDB,ChEMBL |
| IC50 | 200.0 nM | PMID3373486 | BindingDB,ChEMBL |
| IC50 | 2460.0 nM | PMID3351850 | BindingDB,ChEMBL |
| IC50 | 52000.0 nM | PMID3351851 | BindingDB,ChEMBL |
| IC50 | 90000.0 nM | PMID3351851 | BindingDB,ChEMBL |
| Inhibition | 0.93 % | PMID3373486 | ChEMBL |
| Ki | 3.7 nM | PMID2067592 | BindingDB |
| Ki | 5.1 nM | PMID24900277, PMID1554381, PMID23245803 | BindingDB,ChEMBL |
| Ki | 6.24 nM | PMID2995663 | BindingDB,ChEMBL |
| Ki | 6.26 nM | PMID3010074 | BindingDB |
| Ki | 6.3 nM | PMID3336027, PMID8126704, PMID1738138, PMID3385722, PMID9703463, PMID9667957 | BindingDB,ChEMBL |
| Ki | 7.7 nM | PMID11170630 | BindingDB,ChEMBL |
| Ki | 8.2 nM | PMID3373486 | BindingDB |
| Ki | 8.2 nM | PMID3373486, PMID1619615 | BindingDB,ChEMBL |
| Ki | 8.3 nM | PMID1495019 | BindingDB,ChEMBL |
| Ki | 10.0 nM | PMID7739005, PMID7658444, PMID8863798, PMID2795469, PMID1433217, PMID8201607 | BindingDB |
| Ki | 10.3 nM | PMID1433217 | ChEMBL |
| Ki | 10.4 nM | PMID7739005, PMID8863798, PMID8201607, PMID7658444 | ChEMBL |
| Ki | 11.0 nM | PMID1619615 | ChEMBL |
| Ki | 13.0 nM | PMID1732541 | BindingDB |
| Ki | 13.3 nM | PMID1732541, PMID11170643 | BindingDB,ChEMBL |
| Ki | 62.0 nM | PMID7582508 | BindingDB |
| Ki | 63.0 nM | PMID22486652, PMID11784146, PMID7707320, PMID10212124 | BindingDB,ChEMBL |
| Ki | 88.0 nM | PMID7582508 | BindingDB |
| Ki | 170.0 nM | PMID2067592 | BindingDB |
| Ki | 530.0 nM | PMID10212124 | BindingDB,ChEMBL |
| Ki | 650.0 nM | PMID1619615 | BindingDB,ChEMBL |
| Ki | 1257.0 nM | PMID2362269 | BindingDB,ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218