

You can:
| Name | D(3) dopamine receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | DRD3 |
| Synonym | dopaminergic receptor D3 D3R D3 receptor dopamine D3 receptor |
| Disease | Unspecified Emesis; Gastric motility disorder Female sexual dysfunction Male sexual disorders Psychotic disorders [ Show all ] |
| Length | 400 |
| Amino acid sequence | MASLSQLSSHLNYTCGAENSTGASQARPHAYYALSYCALILAIVFGNGLVCMAVLKERALQTTTNYLVVSLAVADLLVATLVMPWVVYLEVTGGVWNFSRICCDVFVTLDVMMCTASILNLCAISIDRYTAVVMPVHYQHGTGQSSCRRVALMITAVWVLAFAVSCPLLFGFNTTGDPTVCSISNPDFVIYSSVVSFYLPFGVTVLVYARIYVVLKQRRRKRILTRQNSQCNSVRPGFPQQTLSPDPAHLELKRYYSICQDTALGGPGFQERGGELKREEKTRNSLSPTIAPKLSLEVRKLSNGRLSTSLKLGPLQPRGVPLREKKATQMVAIVLGAFIVCWLPFFLTHVLNTHCQTCHVSPELYSATTWLGYVNSALNPVIYTTFNIEFRKAFLKILSC |
| UniProt | P35462 |
| Protein Data Bank | 3pbl |
| GPCR-HGmod model | P35462 |
| 3D structure model | This structure is from PDB ID 3pbl. |
| BioLiP | BL0191566, BL0191567 |
| Therapeutic Target Database | T02551 |
| ChEMBL | CHEMBL234 |
| IUPHAR | 216 |
| DrugBank | BE0000581 |
| Name | sulpiride |
|---|---|
| Molecular formula | C15H23N3O4S |
| IUPAC name | N-[(1-ethylpyrrolidin-2-yl)methyl]-2-methoxy-5-sulfamoylbenzamide |
| Molecular weight | 341.426 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 2 |
| XlogP | 0.6 |
| Synonyms | Sulperide Sulpiride, British Pharmacopoeia (BP) Reference Standard Sulpitil Tox21_501050 NCGC00015966-06 [ Show all ] |
| Inchi Key | BGRJTUBHPOOWDU-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C15H23N3O4S/c1-3-18-8-4-5-11(18)10-17-15(19)13-9-12(23(16,20)21)6-7-14(13)22-2/h6-7,9,11H,3-5,8,10H2,1-2H3,(H,17,19)(H2,16,20,21) |
| PubChem CID | 5355 |
| ChEMBL | CHEMBL26 |
| IUPHAR | 5501 |
| BindingDB | 11638 |
| DrugBank | DB00391 |
Structure | 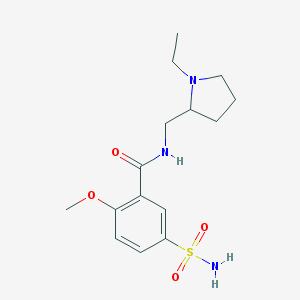 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| N/A | N/A | DrugBank | |
| Emax | -6.83 % | PMID22632094 | ChEMBL |
| IC50 | 66.22 nM | PMID21495689 | BindingDB,ChEMBL |
| IC50 | 169.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| IC50 | 560.0 nM | PMID21495689 | BindingDB,ChEMBL |
| Ki | 7.9 nM | PMID16135699 | PDSP,BindingDB |
| Ki | 8.0 nM | PMID14521403, PMID8301582 | PDSP,BindingDB,ChEMBL |
| Ki | 8.2 nM | PMID16135699 | PDSP,BindingDB |
| Ki | 14.0 - 142.0 nM | PMID7576010 | IUPHAR |
| Ki | 15.0 nM | PMID9686407 | PDSP,BindingDB |
| Ki | 20.0 nM | PMID1586393 | PDSP,BindingDB |
| Ki | 57.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| Ki | 120.0 nM | PMID15689154 | BindingDB,ChEMBL |
| Ki | 480.0 nM | PMID8301582 | PDSP,BindingDB |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218