

You can:
| Name | Histamine H3 receptor |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HRH3 |
| Synonym | HH3R H3R H3 receptor GPCR97 G-protein coupled receptor 97 |
| Disease | Obese insulin-resistant disorders Excessive daytime sleepiness Sleep disorders Schizophrenia Pain [ Show all ] |
| Length | 445 |
| Amino acid sequence | MERAPPDGPLNASGALAGEAAAAGGARGFSAAWTAVLAALMALLIVATVLGNALVMLAFVADSSLRTQNNFFLLNLAISDFLVGAFCIPLYVPYVLTGRWTFGRGLCKLWLVVDYLLCTSSAFNIVLISYDRFLSVTRAVSYRAQQGDTRRAVRKMLLVWVLAFLLYGPAILSWEYLSGGSSIPEGHCYAEFFYNWYFLITASTLEFFTPFLSVTFFNLSIYLNIQRRTRLRLDGAREAAGPEPPPEAQPSPPPPPGCWGCWQKGHGEAMPLHRYGVGEAAVGAEAGEATLGGGGGGGSVASPTSSSGSSSRGTERPRSLKRGSKPSASSASLEKRMKMVSQSFTQRFRLSRDRKVAKSLAVIVSIFGLCWAPYTLLMIIRAACHGHCVPDYWYETSFWLLWANSAVNPVLYPLCHHSFRRAFTKLLCPQKLKIQPHSSLEHCWK |
| UniProt | Q9Y5N1 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9Y5N1 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9Y5N1. |
| BioLiP | N/A |
| Therapeutic Target Database | T64765 |
| ChEMBL | CHEMBL264 |
| IUPHAR | 264 |
| DrugBank | BE0000968 |
| Name | Thioperamide |
|---|---|
| Molecular formula | C15H24N4S |
| IUPAC name | N-cyclohexyl-4-(1H-imidazol-5-yl)piperidine-1-carbothioamide |
| Molecular weight | 292.445 |
| Hydrogen bond acceptor | 2 |
| Hydrogen bond donor | 2 |
| XlogP | 2.4 |
| Synonyms | AKOS024458593 CHEBI:81391 KS-000001KM N-Cyclohexy-4-(imidazol-4-yl)-1-piperidinecarbothioamide NCGC00015988-04 [ Show all ] |
| Inchi Key | QKDDJDBFONZGBW-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C15H24N4S/c20-15(18-13-4-2-1-3-5-13)19-8-6-12(7-9-19)14-10-16-11-17-14/h10-13H,1-9H2,(H,16,17)(H,18,20) |
| PubChem CID | 3035905 |
| ChEMBL | CHEMBL260374 |
| IUPHAR | 1267 |
| BindingDB | 22914 |
| DrugBank | N/A |
Structure | 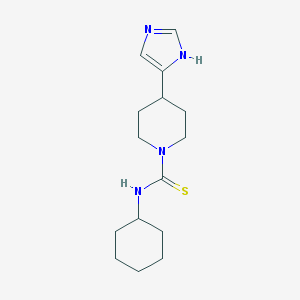 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Activity | -0.71 - | PMID19317445 | ChEMBL |
| EC50 | 39.81 nM | PMID21348462 | BindingDB,ChEMBL |
| EC50 | 97.72 nM | PMID19317445, PMID19791743 | BindingDB,ChEMBL |
| EC50 | 98.0 nM | PMID19317445 | BindingDB |
| EC50 | 100.0 nM | MedChemComm, (2010) 1:1:39 | ChEMBL |
| IC50 | 16.0 nM | PMID16942032 | BindingDB,ChEMBL |
| IC50 | 120.0 nM | PMID23978359 | ChEMBL |
| IC50 | 517.5 nM | PMID22019465 | ChEMBL |
| IC50 | 518.0 nM | PMID22019465 | BindingDB |
| Inhibition | 98.0 % | PMID20397706, PMID16942032 | ChEMBL |
| Intrinsic activity | -0.59 - | PMID27007611 | ChEMBL |
| Kb | 78.0 nM | PMID27007611 | ChEMBL |
| Kd | 43.65 nM | PMID15771465 | BindingDB,ChEMBL |
| Kd | 63.1 nM | PMID19577344 | ChEMBL |
| Ki | <10000.0 nM | PMID10347254 | PDSP,BindingDB |
| Ki | 14.0 nM | PMID15139761, PMID15634025 | BindingDB,ChEMBL |
| Ki | 19.9 - 79.4 nM | PMID12606603, PMID15294456, PMID10869375, PMID11284713, PMID11714875, PMID11090094, PMID12393057 | IUPHAR |
| Ki | 25.0 nM | PMID12954048, PMID11179434 | PDSP,BindingDB,ChEMBL |
| Ki | 28.84 nM | PMID15771465 | BindingDB,ChEMBL |
| Ki | 29.51 nM | PMID21348462 | BindingDB,ChEMBL |
| Ki | 31.62 nM | PMID19524331 | BindingDB,ChEMBL |
| Ki | 39.81 nM | PMID15466448 | BindingDB |
| Ki | 39.8107 nM | PMID15466448 | PDSP |
| Ki | 50.0 nM | PMID15947036 | BindingDB |
| Ki | 50.12 nM | PMID19414267, PMID21062081 | BindingDB,ChEMBL |
| Ki | 51.1 nM | PMID20397706, PMID16942032 | BindingDB,ChEMBL |
| Ki | 58.0 nM | PMID10869375 | PDSP,BindingDB |
| Ki | 60.0 nM | PMID19846299, PMID12672253, PMID21498080, PMID11294398 | BindingDB,ChEMBL |
| Ki | 61.3 nM | PMID15033391 | PDSP,BindingDB |
| Ki | 63.0 nM | PMID19577344 | BindingDB,ChEMBL |
| Ki | 63.1 nM | PMID19773175 | ChEMBL |
| Ki | 66.0693 nM | PMID11714875, PMID12606603 | PDSP |
| Ki | 66.07 nM | PMID11714875, PMID12606603, PMID18683917 | BindingDB,ChEMBL |
| Ki | 70.79 nM | PMID24650714 | ChEMBL |
| Ki | 71.0 nM | PMID24650714 | BindingDB |
| Ki | 72.0 nM | PMID15634000, PMID18683917 | BindingDB,ChEMBL |
| Ki | 72.44 nM | PMID15634000, PMID12606603, PMID18683917 | BindingDB,ChEMBL |
| Ki | 72.4436 nM | PMID12606603 | PDSP |
| Ki | 72.6 nM | PMID15033391 | BindingDB |
| Ki | 158.49 nM | MedChemComm, (2010) 1:1:39 | ChEMBL |
| Ki | 602.56 nM | PMID23891186 | ChEMBL |
| pKb | 6.1 - | PMID18683917 | ChEMBL |
| pKb | 6.82 - | PMID15634000, PMID18683917 | ChEMBL |
| pKb | 7.01 - | MedChemComm, (2014) 5:1:72 | ChEMBL |
| pKb | 7.39 - | PMID18683917 | ChEMBL |
| Ratio | 9.0 - | PMID11294398 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218