

You can:
| Name | Mu-type opioid receptor |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Oprm1 |
| Synonym | opioid receptor, mu 1 opioid receptor OP3 Mu receptor MOR-1 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 398 |
| Amino acid sequence | MDSSAGPGNISDCSDPLAPASCSPAPGSWLNLSHVDGNQSDPCGPNRTGLGGSHSLCPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALATSTLPFQSVNYLMGTWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKYRQGSIDCTLTFSHPTWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMLSGSKEKDRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALITIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCFREFCIPTSSTIEQQNSARIRQNTREHPSTANTVDRTNHQLENLEAETAPLP |
| UniProt | P42866 |
| Protein Data Bank | 4dkl, 5c1m, 6dde, 6ddf |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 4dkl. |
| BioLiP | BL0416752, BL0416751, BL0321492, BL0224753, BL0224754, BL0224755,BL0224756, BL0321491 |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL2858 |
| IUPHAR | 319 |
| DrugBank | N/A |
| Name | morphine |
|---|---|
| Molecular formula | C17H19NO3 |
| IUPAC name | (4R,4aR,7S,7aR,12bS)-3-methyl-2,4,4a,7,7a,13-hexahydro-1H-4,12-methanobenzofuro[3,2-e]isoquinoline-7,9-diol |
| Molecular weight | 285.343 |
| Hydrogen bond acceptor | 4 |
| Hydrogen bond donor | 2 |
| XlogP | 0.8 |
| Synonyms | 4-methyl-(1S,5R,13R,14S,17R)-12-oxa-4-azapentacyclo[9.6.1.01,13.05,17.07,18]octadeca-7(18),8,10,15-tetraene-10,14-diol Morphin 4-methyl-(1S,5R,13R,14S,17R)-12-oxa-4-azapentacyclo[9.6.1.01,13.05,17.07,18]octadeca-7(18),8,10,15-tetraene-10,14-diolMorphine 6-tert-Butyl-3-methyl-1,2,3,4,5,6-hexahydro-2,6-methano-benzo[d]azocine Morphine solution, 1.0 mg/mL in methanol, ampule of 1 mL, certified reference material [ Show all ] |
| Inchi Key | BQJCRHHNABKAKU-KBQPJGBKSA-N |
| Inchi ID | InChI=1S/C17H19NO3/c1-18-7-6-17-10-3-5-13(20)16(17)21-15-12(19)4-2-9(14(15)17)8-11(10)18/h2-5,10-11,13,16,19-20H,6-8H2,1H3/t10-,11+,13-,16-,17-/m0/s1 |
| PubChem CID | 5288826 |
| ChEMBL | CHEMBL70 |
| IUPHAR | 1627 |
| BindingDB | 50000092 |
| DrugBank | DB00295 |
Structure | 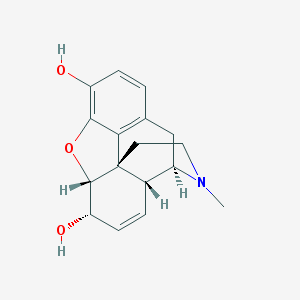 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Activity | 1.0 nM | PMID25783191 | ChEMBL |
| EC50 | 30.0 nM | PMID23880538 | ChEMBL |
| EC50 | 85.5 nM | PMID20413312 | BindingDB,ChEMBL |
| EC50 | 120.0 nM | PMID25783191 | BindingDB,ChEMBL |
| EC50 | 395.0 nM | PMID17962026 | BindingDB,ChEMBL |
| Emax | 24.41 % | PMID20413312 | ChEMBL |
| Emax | 106.0 % | PMID25783191 | ChEMBL |
| IC50 | 1.27 nM | PMID18337104 | BindingDB,ChEMBL |
| IC50 | 1.271 nM | PMID18990576 | ChEMBL |
| IC50 | 1.3 nM | PMID18990576 | BindingDB |
| IC50 | 1420.0 nM | PMID6090663 | BindingDB,ChEMBL |
| Inhibition | 97.0 % | PMID23880538 | ChEMBL |
| Ke | 3.29 nM | PMID18829333 | ChEMBL |
| Ki | 0.45 nM | PMID9694962 | BindingDB |
| Ki | 1.2 nM | PMID25562563 | BindingDB |
| Ki | 1.68 nM | PMID9694962 | BindingDB |
| Ki | 1.8 nM | PMID15482911 | BindingDB,ChEMBL |
| Ratio | 0.02 - | PMID15482911 | ChEMBL |
| Ratio | 3.0 - | PMID8394935 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218