

You can:
| Name | 5-hydroxytryptamine receptor 6 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | HTR6 |
| Synonym | 5-HT6 receptor 5-hydroxytryptamine (serotonin) receptor 6, G protein-coupled 5-HT-6 serotonin receptor 6 ST-B17 [ Show all ] |
| Disease | Schizophrenia Obesity Neurological disease Neurodegenerative disease Emesis [ Show all ] |
| Length | 440 |
| Amino acid sequence | MVPEPGPTANSTPAWGAGPPSAPGGSGWVAAALCVVIALTAAANSLLIALICTQPALRNTSNFFLVSLFTSDLMVGLVVMPPAMLNALYGRWVLARGLCLLWTAFDVMCCSASILNLCLISLDRYLLILSPLRYKLRMTPLRALALVLGAWSLAALASFLPLLLGWHELGHARPPVPGQCRLLASLPFVLVASGLTFFLPSGAICFTYCRILLAARKQAVQVASLTTGMASQASETLQVPRTPRPGVESADSRRLATKHSRKALKASLTLGILLGMFFVTWLPFFVANIVQAVCDCISPGLFDVLTWLGYCNSTMNPIIYPLFMRDFKRALGRFLPCPRCPRERQASLASPSLRTSHSGPRPGLSLQQVLPLPLPPDSDSDSDAGSGGSSGLRLTAQLLLPGEATQDPPLPTRAAAAVNFFNIDPAEPELRPHPLGIPTN |
| UniProt | P50406 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | P50406 |
| 3D structure model | This predicted structure model is from GPCR-EXP P50406. |
| BioLiP | N/A |
| Therapeutic Target Database | T16691 |
| ChEMBL | CHEMBL3371 |
| IUPHAR | 11 |
| DrugBank | BE0000945 |
| Name | chlorpromazine |
|---|---|
| Molecular formula | C17H19ClN2S |
| IUPAC name | 3-(2-chlorophenothiazin-10-yl)-N,N-dimethylpropan-1-amine |
| Molecular weight | 318.863 |
| Hydrogen bond acceptor | 3 |
| Hydrogen bond donor | 0 |
| XlogP | 5.2 |
| Synonyms | 10H-Phenothiazine-10-propanamine, 2-chloro-N,N-dimethyl-, radical ion(1+) Phenothiazine, 2-chloro-10-(3-(dimethylamino)propyl)- Chlorpromazine [USAN:INN:BAN] 2-Chloropromazine Prestwick3_000064 [ Show all ] |
| Inchi Key | ZPEIMTDSQAKGNT-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C17H19ClN2S/c1-19(2)10-5-11-20-14-6-3-4-7-16(14)21-17-9-8-13(18)12-15(17)20/h3-4,6-9,12H,5,10-11H2,1-2H3 |
| PubChem CID | 2726 |
| ChEMBL | CHEMBL71 |
| IUPHAR | 83 |
| BindingDB | 50001888 |
| DrugBank | DB00477 |
Structure | 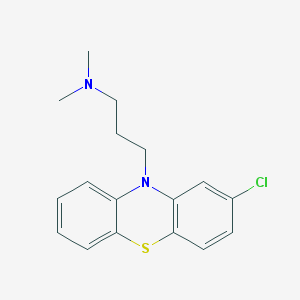 |
| Lipinski's druglikeness | This ligand has a partition coefficient log P greater than 5. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| N/A | N/A | DrugBank | |
| IC50 | 57.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| Ki | 4.0 nM | PMID12825922 | BindingDB,ChEMBL |
| Ki | 12.0 nM | http://pdsp.med.unc.edu/pdsp.php | PDSP |
| Ki | 15.8489 - 19.9526 nM | PMID8522988, PMID15821958 | IUPHAR |
| Ki | 19.0 nM | PMID8522988 | PDSP,BindingDB |
| Ki | 20.1 nM | PMID12629531 | PDSP,BindingDB |
| Ki | 26.0 nM | DrugMatrix in vitro pharmacology data | ChEMBL |
| Ki | >50.0 nM | PMID15771424 | ChEMBL |
| Ki | 50.12 nM | PMID17880057 | ChEMBL |
| Ki | 60.0 nM | PMID24365159 | ChEMBL |
| Ki | 60.26 nM | PMID24365159 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218