

You can:
| Name | Neuromedin-U receptor 1 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | NMUR1 |
| Synonym | NMU1 receptor NmU-R1 GPR66 G-protein coupled receptor FM-3 G-protein coupled receptor 66 [ Show all ] |
| Disease | N/A |
| Length | 426 |
| Amino acid sequence | MTPLCLNCSVLPGDLYPGGARNPMACNGSAARGHFDPEDLNLTDEALRLKYLGPQQTELFMPICATYLLIFVVGAVGNGLTCLVILRHKAMRTPTNYYLFSLAVSDLLVLLVGLPLELYEMWHNYPFLLGVGGCYFRTLLFEMVCLASVLNVTALSVERYVAVVHPLQARSMVTRAHVRRVLGAVWGLAMLCSLPNTSLHGIRQLHVPCRGPVPDSAVCMLVRPRALYNMVVQTTALLFFCLPMAIMSVLYLLIGLRLRRERLLLMQEAKGRGSAAARSRYTCRLQQHDRGRRQVTKMLFVLVVVFGICWAPFHADRVMWSVVSQWTDGLHLAFQHVHVISGIFFYLGSAANPVLYSLMSSRFRETFQEALCLGACCHRLRPRHSSHSLSRMTTGSTLCDVGSLGSWVHPLAGNDGPEAQQETDPS |
| UniProt | Q9HB89 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | Q9HB89 |
| 3D structure model | This predicted structure model is from GPCR-EXP Q9HB89. |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1075178 |
| IUPHAR | 298 |
| DrugBank | N/A |
| Name | CHEMBL3759206 |
|---|---|
| Molecular formula | C69H114N18O11 |
| IUPAC name | (2S)-2-[[(2S)-5-(diaminomethylideneamino)-2-[[(2S)-1-[(2S)-5-(diaminomethylideneamino)-2-[[(2S)-2-[[(2S)-2-[[(2S)-2-[[2-[[2-[[2-(hexadecylamino)-2-oxoethyl]amino]acetyl]amino]-2-methylpropanoyl]amino]-3-phenylpropanoyl]amino]-4-methylpentanoyl]amino]-3-phenylpropanoyl]amino]pentanoyl]pyrrolidine-2-carbonyl]amino]pentanoyl]amino]butanediamide |
| Molecular weight | 1371.79 |
| Hydrogen bond acceptor | 14 |
| Hydrogen bond donor | 15 |
| XlogP | 4.9 |
| Synonyms | SCHEMBL18480625 |
| Inchi Key | FMLNZTDPRUXRKS-DKKXDTQASA-N |
| Inchi ID | InChI=1S/C69H114N18O11/c1-6-7-8-9-10-11-12-13-14-15-16-17-18-25-36-77-57(89)44-76-45-58(90)86-69(4,5)66(98)85-54(42-48-31-23-20-24-32-48)63(95)83-52(40-46(2)3)61(93)84-53(41-47-29-21-19-22-30-47)62(94)81-50(34-27-38-79-68(74)75)65(97)87-39-28-35-55(87)64(96)80-49(33-26-37-78-67(72)73)60(92)82-51(59(71)91)43-56(70)88/h19-24,29-32,46,49-55,76H,6-18,25-28,33-45H2,1-5H3,(H2,70,88)(H2,71,91)(H,77,89)(H,80,96)(H,81,94)(H,82,92)(H,83,95)(H,84,93)(H,85,98)(H,86,90)(H4,72,73,78)(H4,74,75,79)/t49-,50-,51-,52-,53-,54-,55-/m0/s1 |
| PubChem CID | 126558991 |
| ChEMBL | CHEMBL3759206 |
| IUPHAR | N/A |
| BindingDB | N/A |
| DrugBank | N/A |
Structure | 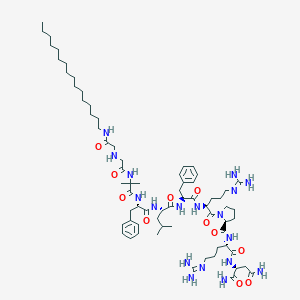 |
| Lipinski's druglikeness | This ligand has more than 5 hydrogen bond donor. This ligand has more than 10 hydrogen bond acceptor. This ligand is heavier than 500 daltons. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| EC50 | 12.3 nM | PMID26204509 | ChEMBL |
| Emax | 55.7 % | PMID26204509 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218