

You can:
| Name | 5-hydroxytryptamine receptor 2A |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Htr2a |
| Synonym | serotonin 5HT-2 receptor 5Ht-2 'D' receptor 5-hydroxytryptamine (serotonin) receptor 2A, G protein-coupled 5-HT2A receptor [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 471 |
| Amino acid sequence | MEILCEDNISLSSIPNSLMQLGDGPRLYHNDFNSRDANTSEASNWTIDAENRTNLSCEGYLPPTCLSILHLQEKNWSALLTTVVIILTIAGNILVIMAVSLEKKLQNATNYFLMSLAIADMLLGFLVMPVSMLTILYGYRWPLPSKLCAIWIYLDVLFSTASIMHLCAISLDRYVAIQNPIHHSRFNSRTKAFLKIIAVWTISVGISMPIPVFGLQDDSKVFKEGSCLLADDNFVLIGSFVAFFIPLTIMVITYFLTIKSLQKEATLCVSDLSTRAKLASFSFLPQSSLSSEKLFQRSIHREPGSYAGRRTMQSISNEQKACKVLGIVFFLFVVMWCPFFITNIMAVICKESCNENVIGALLNVFVWIGYLSSAVNPLVYTLFNKTYRSAFSRYIQCQYKENRKPLQLILVNTIPALAYKSSQLQVGQKKNSQEDAEQTVDDCSMVTLGKQQSEENCTDNIETVNEKVSCV |
| UniProt | P14842 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL322 |
| IUPHAR | 6 |
| DrugBank | N/A |
| Name | clozapine |
|---|---|
| Molecular formula | C18H19ClN4 |
| IUPAC name | 3-chloro-6-(4-methylpiperazin-1-yl)-11H-benzo[b][1,4]benzodiazepine |
| Molecular weight | 326.828 |
| Hydrogen bond acceptor | 3 |
| Hydrogen bond donor | 1 |
| XlogP | 3.1 |
| Synonyms | I06-0047 Clorazil (E)-8-chloro-11-(4-methylpiperazin-1-yl)-5H-dibenzo[b,e][1,4]diazepine Clozapine for peak identification, European Pharmacopoeia (EP) Reference Standard 5786-21-0 [ Show all ] |
| Inchi Key | QZUDBNBUXVUHMW-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C18H19ClN4/c1-22-8-10-23(11-9-22)18-14-4-2-3-5-15(14)20-16-7-6-13(19)12-17(16)21-18/h2-7,12,20H,8-11H2,1H3 |
| PubChem CID | 135398737 |
| ChEMBL | CHEMBL42 |
| IUPHAR | 38 |
| BindingDB | 50001884 |
| DrugBank | DB00363 |
Structure | 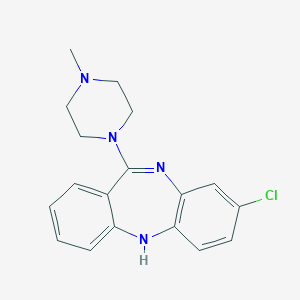 |
| Lipinski's druglikeness | This ligand satisfies Lipinski's rule of five. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Affinity ratio | 5.5 - | PMID8568801 | ChEMBL |
| IC50 | 7.8 nM | PMID9406603, PMID7473566 | BindingDB,ChEMBL |
| IC50 | 12.0 nM | PMID10377229 | BindingDB,ChEMBL |
| IC50 | 13.0 nM | PMID9876110 | BindingDB,ChEMBL |
| IC50 | 40.7 nM | PMID12361392 | BindingDB,ChEMBL |
| IC50 | 72.0 nM | PMID7914536 | ChEMBL |
| IC50 | 72.0 nM | PMID7914536 | BindingDB |
| Kd | 0.6918 nM | PMID11754579 | ChEMBL |
| Kd | 0.692 nM | PMID11754579 | BindingDB |
| Kd | 2.399 nM | PMID9934473 | ChEMBL |
| Ki | 1.25893 nM | PMID9655845 | IUPHAR |
| Ki | 1.585 nM | PMID10821703 | ChEMBL |
| Ki | 5.0 nM | N/A | BindingDB |
| Ki | 5.012 nM | Bioorg. Med. Chem. Lett., (1997) 7:7:913, PMID10425088 | ChEMBL |
| Ki | 6.05 nM | PMID18603331 | BindingDB,ChEMBL |
| Ki | 6.7 nM | PMID12408724 | BindingDB,ChEMBL |
| Ki | 7.586 nM | PMID9934473, PMID11754579 | ChEMBL |
| Ki | 7.6 nM | PMID9934473 | BindingDB |
| Ki | 8.0 nM | PMID7861418, PMID8064797 | BindingDB,ChEMBL |
| Ki | 8.9 nM | PMID21816515 | BindingDB,ChEMBL |
| Ki | 9.0 nM | PMID10715164 | BindingDB,ChEMBL |
| Ki | 9.4 nM | PMID9876110 | BindingDB,ChEMBL |
| Ki | 10.0 nM | PMID15771414, PMID11784139, PMID19072656, PMID20481570, PMID14695828 | BindingDB,ChEMBL |
| Ki | 11.6 nM | PMID23675993 | ChEMBL |
| Ki | 12.9 nM | PMID24487191, MedChemComm, (2015) 6:5:831 | ChEMBL |
| Ki | 13.18 nM | PMID17870534 | ChEMBL |
| Ki | 14.0 nM | PMID20153652 | BindingDB,ChEMBL |
| Ki | 14.5 nM | PMID26483200 | ChEMBL |
| Ki | 15.0 nM | PMID26483200 | BindingDB |
| Ki | 15.14 nM | PMID8568801 | ChEMBL |
| Ki | 17.0 nM | PMID8709107 | BindingDB,ChEMBL |
| Ki | 20.0 nM | PMID23353740 | BindingDB,ChEMBL |
| Ki | 21.0 nM | PMID23792350 | ChEMBL |
| Ki | 27.54 nM | PMID11101359 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218