

You can:
| Name | C-C chemokine receptor type 5 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | CCR5 |
| Synonym | CD195 chemokine (C-C motif) receptor 5 (gene/pseudogene) CHEMR13 CCR5 CCR-5 [ Show all ] |
| Disease | Human immunodeficiency virus infection |
| Length | 352 |
| Amino acid sequence | MDYQVSSPIYDINYYTSEPCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINCKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPGIIFTRSQKEGLHYTCSSHFPYSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRCRNEKKRHRAVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEKFRNYLLVFFQKHIAKRFCKCCSIFQQEAPERASSVYTRSTGEQEISVGL |
| UniProt | P51681 |
| Protein Data Bank | 4mbs, 6aky, 6akx, 5uiw |
| GPCR-HGmod model | P51681 |
| 3D structure model | This structure is from PDB ID 4mbs. |
| BioLiP | BL0430746, BL0430745, BL0430742,BL0430744, BL0430741,BL0430743, BL0382816, BL0256313,BL0256315, BL0256312,BL0256314 |
| Therapeutic Target Database | T72171 |
| ChEMBL | CHEMBL274 |
| IUPHAR | 62 |
| DrugBank | BE0000911 |
| Name | CHEMBL263529 |
|---|---|
| Molecular formula | C37H51FN4O2 |
| IUPAC name | (2R)-2-[(3S,4S)-3-[[4-[2-ethyl-5-[(4-propan-2-ylphenyl)methyl]pyrazol-3-yl]piperidin-1-yl]methyl]-4-(3-fluorophenyl)pyrrolidin-1-yl]-3,3-dimethylbutanoic acid |
| Molecular weight | 602.839 |
| Hydrogen bond acceptor | 6 |
| Hydrogen bond donor | 1 |
| XlogP | 5.2 |
| Synonyms | (2R)-2-[(3S,4S)-3-({4-[1-Ethyl-3-(4-isopropylbenzyl)-1H-pyrazol-5-yl]piperidin-1-yl}methyl)-4-(3-fluorophenyl)pyrrolidin-1-yl]-3,3-dimethylbutanoic acid (R)-2-[(2S,3S)-3-{4-[2-Ethyl-5-(4-isopropyl-benzyl)-2H-pyrazol-3-yl]-piperidin-1-ylmethyl}-4-(3-fluoro-phenyl)-pyrrolidin-1-yl]-3,3-dimethyl-butyric acid BDBM50141968 (2R)-2-[(3S,4S)-3-[[4-[2-ethyl-5-[(4-isopropylphenyl)methyl]pyrazol-3-yl]-1-piperidyl]methyl]-4-(3-fluorophenyl)pyrrolidin-1-yl]-3,3-dimethyl-butanoic acid (2R)-2-[(3S,4S)-3-[[4-[2-ethyl-5-[(4-propan-2-ylphenyl)methyl]pyrazol-3-yl]piperidin-1-yl]methyl]-4-(3-fluorophenyl)pyrrolidin-1-yl]-3,3-dimethylbutanoic acid [ Show all ] |
| Inchi Key | ALNIPKRVLOOYCX-LYEZUSGXSA-N |
| Inchi ID | InChI=1S/C37H51FN4O2/c1-7-42-34(21-32(39-42)19-26-11-13-27(14-12-26)25(2)3)28-15-17-40(18-16-28)22-30-23-41(35(36(43)44)37(4,5)6)24-33(30)29-9-8-10-31(38)20-29/h8-14,20-21,25,28,30,33,35H,7,15-19,22-24H2,1-6H3,(H,43,44)/t30-,33+,35-/m0/s1 |
| PubChem CID | 516291 |
| ChEMBL | CHEMBL263529 |
| IUPHAR | N/A |
| BindingDB | 50141968 |
| DrugBank | N/A |
Structure | 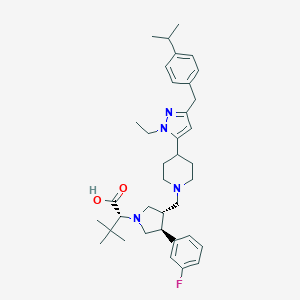 |
| Lipinski's druglikeness | This ligand is heavier than 500 daltons. This ligand has a partition coefficient log P greater than 5. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| IC50 | 0.6 nM | PMID15012999 | BindingDB |
| IC50 | 0.6 nM | PMID15012999 | ChEMBL |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218