

You can:
| Name | Neurotensin receptor type 1 |
|---|---|
| Species | Rattus norvegicus (Rat) |
| Gene | Ntsr1 |
| Synonym | NTS1 receptor NTRH NTR1 NTR NT-R-1 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 424 |
| Amino acid sequence | MHLNSSVPQGTPGEPDAQPFSGPQSEMEATFLALSLSNGSGNTSESDTAGPNSDLDVNTDIYSKVLVTAIYLALFVVGTVGNSVTAFTLARKKSLQSLQSTVHYHLGSLALSDLLILLLAMPVELYNFIWVHHPWAFGDAGCRGYYFLRDACTYATALNVASLSVERYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNRSGDGTHPGGLVCTPIVDTATVKVVIQVNTFMSFLFPMLVISILNTVIANKLTVMVHQAAEQGRVCTVGTHNGLEHSTFNMTIEPGRVQALRHGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDEQWTTFLFDFYHYFYMLTNALFYVSSAINPILYNLVSANFRQVFLSTLACLCPGWRHRRKKRPTFSRKPNSMSSNHAFSTSATRETLY |
| UniProt | P20789 |
| Protein Data Bank | 3zev, 4buo, 4bv0, 4bwb, 5t04 |
| GPCR-HGmod model | N/A |
| 3D structure model | This structure is from PDB ID 3zev. |
| BioLiP | BL0365097, BL0267346,BL0267350, BL0267315,BL0267316, BL0267314,BL0267317, BL0267354,BL0267355, BL0267356,BL0267357, BL0365096, BL0267347,BL0267348,BL0267349, |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL3027 |
| IUPHAR | 309 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 18392 | 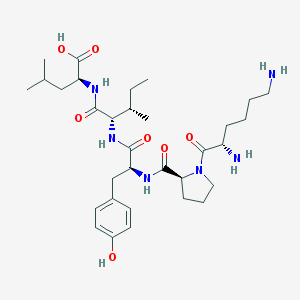 CHEMBL8454 CHEMBL8454 | C32H52N6O7 | 632.803 | 9 / 7 | -1.8 | No |
| 23048 | 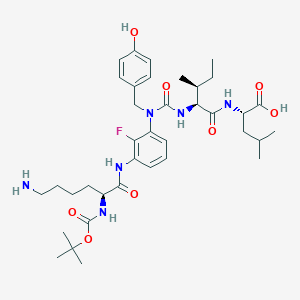 CHEMBL1794000 CHEMBL1794000 | C37H55FN6O8 | 730.879 | 10 / 7 | 2.5 | No |
| 23300 | 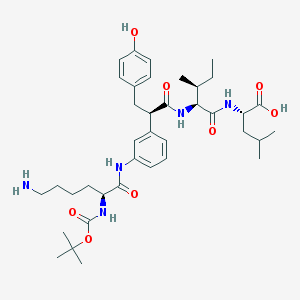 CHEMBL1794002 CHEMBL1794002 | C38H57N5O8 | 711.901 | 9 / 7 | 3.0 | No |
| 553401 | 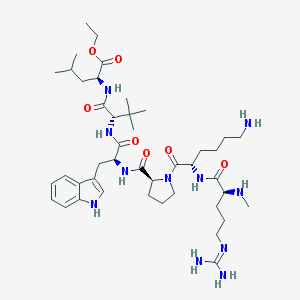 EISAI-1 EISAI-1 | C43H71N11O7 | 854.111 | 10 / 9 | 2.5 | No |
| 28443 | 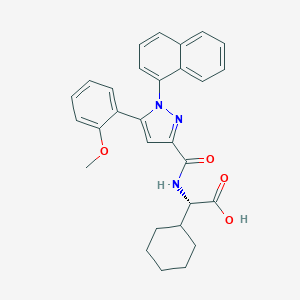 CHEMBL3290100 CHEMBL3290100 | C29H29N3O4 | 483.568 | 5 / 2 | 6.4 | No |
| 32260 | 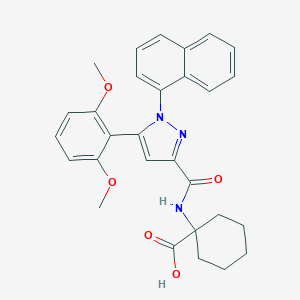 CHEMBL3290101 CHEMBL3290101 | C29H29N3O5 | 499.567 | 6 / 2 | 5.6 | No |
| 34907 | 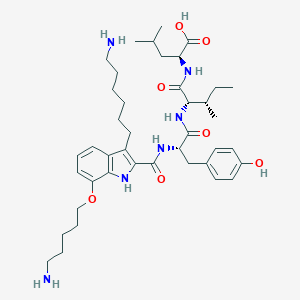 CHEMBL21521 CHEMBL21521 | C41H62N6O7 | 750.982 | 9 / 8 | 4.0 | No |
| 36340 | 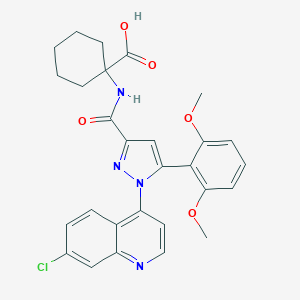 CHEMBL3290096 CHEMBL3290096 | C28H27ClN4O5 | 534.997 | 7 / 2 | 5.2 | No |
| 36560 | 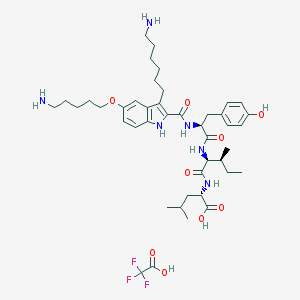 CHEMBL278301 CHEMBL278301 | C43H63F3N6O9 | 865.005 | 14 / 9 | N/A | No |
| 48964 | 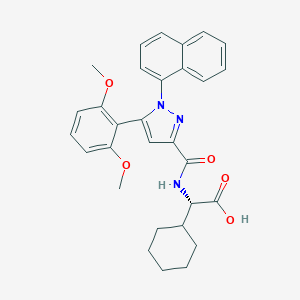 CHEMBL3290099 CHEMBL3290099 | C30H31N3O5 | 513.594 | 6 / 2 | 6.4 | No |
| 50568 |  CHEMBL1790472 CHEMBL1790472 | C37H60N6O8 | 716.921 | 9 / 7 | -0.1 | No |
| 55081 | 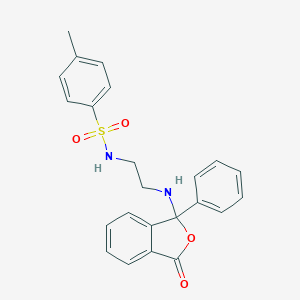 CHEMBL38243 CHEMBL38243 | C23H22N2O4S | 422.499 | 6 / 2 | 3.6 | Yes |
| 61348 | 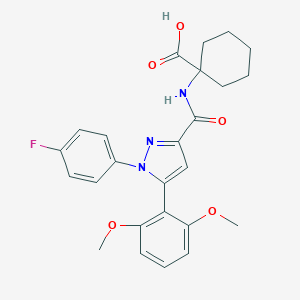 CHEMBL3290105 CHEMBL3290105 | C25H26FN3O5 | 467.497 | 7 / 2 | 4.4 | Yes |
| 61637 | 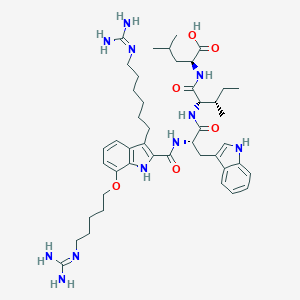 CHEMBL21892 CHEMBL21892 | C45H67N11O6 | 858.102 | 8 / 10 | 5.5 | No |
| 65553 | 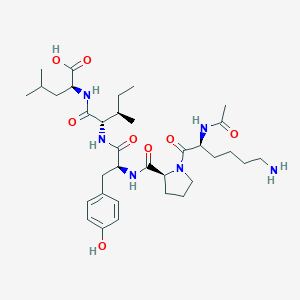 CHEMBL1790473 CHEMBL1790473 | C34H54N6O8 | 674.84 | 9 / 7 | -1.5 | No |
| 66204 | 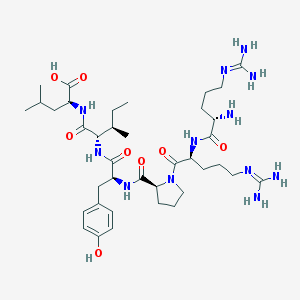 CHEMBL342252 CHEMBL342252 | C38H64N12O8 | 817.006 | 11 / 11 | -4.0 | No |
| 471090 | 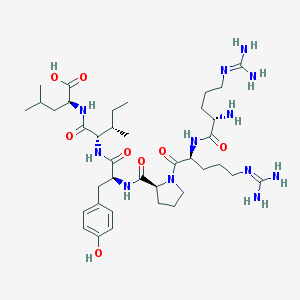 neurotensin (8-13) neurotensin (8-13) | C38H64N12O8 | 817.006 | 11 / 11 | -4.0 | No |
| 67931 |  CHEMBL1790465 CHEMBL1790465 | C45H60N6O8 | 813.009 | 9 / 7 | 3.5 | No |
| 68131 |  CHEMBL1172376 CHEMBL1172376 | C41H70ClN9O8 | 852.516 | 10 / 8 | N/A | No |
| 70331 | 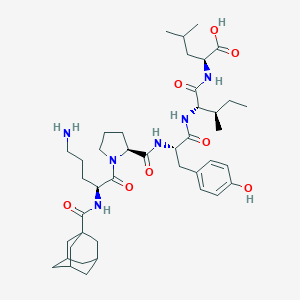 CHEMBL1790463 CHEMBL1790463 | C42H64N6O8 | 781.008 | 9 / 7 | 2.4 | No |
| 444520 | 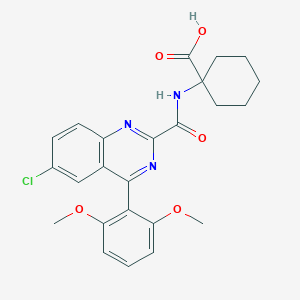 CHEMBL3356853 CHEMBL3356853 | C24H24ClN3O5 | 469.922 | 7 / 2 | 4.6 | Yes |
| 72065 | 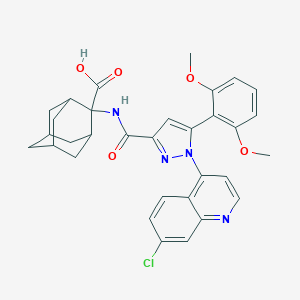 Meclinertant Meclinertant | C32H31ClN4O5 | 587.073 | 7 / 2 | 5.8 | No |
| 553629 | 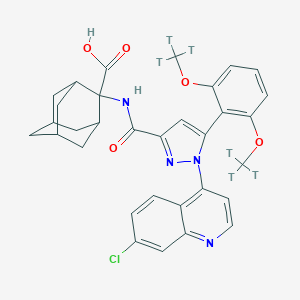 [3H]reminertant [3H]reminertant | C32H31ClN4O5 | 599.121 | 7 / 2 | 5.8 | No |
| 72431 | 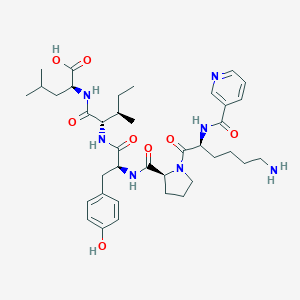 CHEMBL1790466 CHEMBL1790466 | C38H55N7O8 | 737.899 | 10 / 7 | 0.8 | No |
| 79936 | 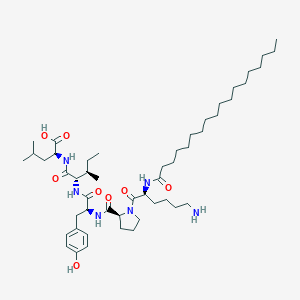 CHEMBL1790476 CHEMBL1790476 | C50H86N6O8 | 899.272 | 9 / 7 | 8.6 | No |
| 82731 | 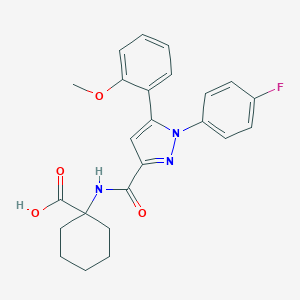 CHEMBL3290106 CHEMBL3290106 | C24H24FN3O4 | 437.471 | 6 / 2 | 4.4 | Yes |
| 474853 | 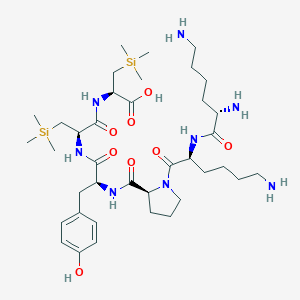 CHEMBL3622803 CHEMBL3622803 | C38H68N8O8Si2 | 821.18 | 11 / 9 | N/A | No |
| 113661 | 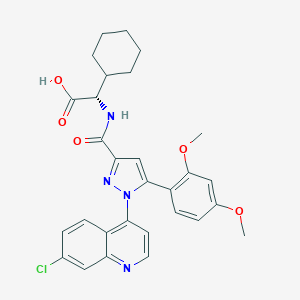 CHEMBL3290091 CHEMBL3290091 | C29H29ClN4O5 | 549.024 | 7 / 2 | 6.0 | No |
| 118103 |  CHEMBL427656 CHEMBL427656 | C36H58N6O9 | 718.893 | 10 / 7 | -0.5 | No |
| 477518 | 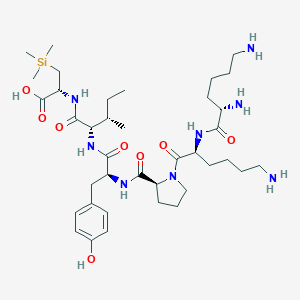 CHEMBL3622802 CHEMBL3622802 | C38H66N8O8Si | 791.079 | 11 / 9 | N/A | No |
| 120045 | 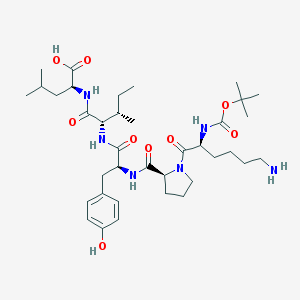 CHEMBL8857 CHEMBL8857 | C37H60N6O9 | 732.92 | 10 / 7 | 1.6 | No |
| 129830 | 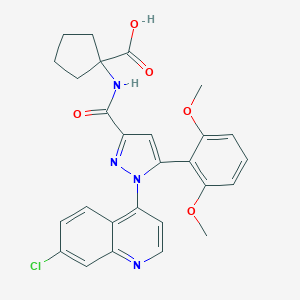 CHEMBL3290097 CHEMBL3290097 | C27H25ClN4O5 | 520.97 | 7 / 2 | 4.7 | No |
| 138504 | 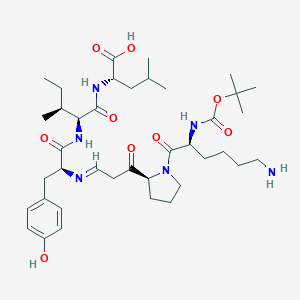 BDBM50281810 BDBM50281810 | C39H62N6O9 | 758.958 | 11 / 6 | 1.9 | No |
| 145858 | 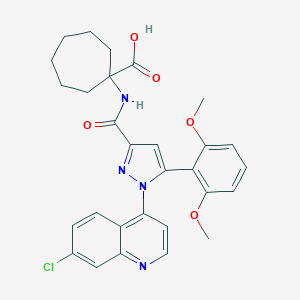 CHEMBL3290095 CHEMBL3290095 | C29H29ClN4O5 | 549.024 | 7 / 2 | 5.7 | No |
| 481836 | 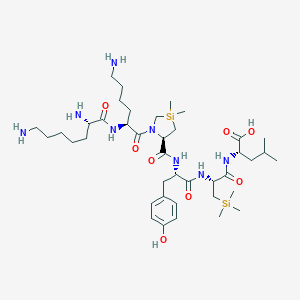 CHEMBL3622804 CHEMBL3622804 | C40H72N8O8Si2 | 849.234 | 11 / 9 | N/A | No |
| 156158 |  CHEMBL1793999 CHEMBL1793999 | C37H55FN6O7 | 714.88 | 9 / 6 | 2.9 | No |
| 482998 | 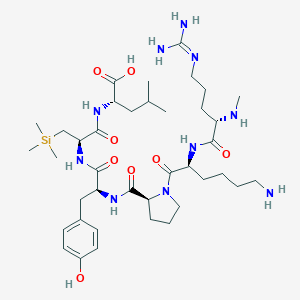 CHEMBL3622805 CHEMBL3622805 | C39H68N10O8Si | 833.12 | 11 / 10 | N/A | No |
| 173966 | 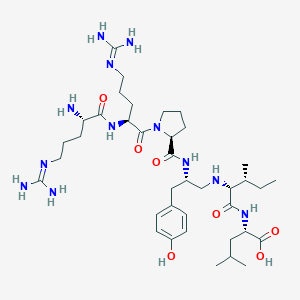 CHEMBL1793838 CHEMBL1793838 | C38H66N12O7 | 803.023 | 11 / 11 | -2.2 | No |
| 180784 | 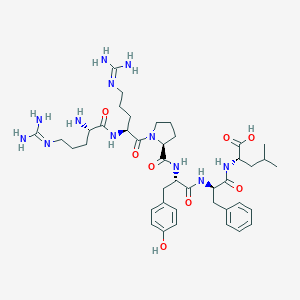 CHEMBL2371276 CHEMBL2371276 | C41H62N12O8 | 851.023 | 11 / 11 | -3.7 | No |
| 184075 | 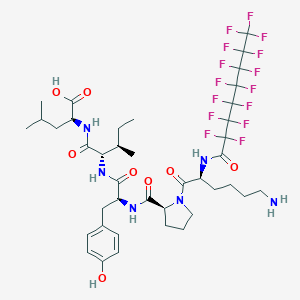 CHEMBL1790469 CHEMBL1790469 | C40H51F15N6O8 | 1028.86 | 24 / 7 | 5.3 | No |
| 196974 |  CHEMBL270332 CHEMBL270332 | C76H124N16O14 | 1485.93 | 18 / 16 | 4.4 | No |
| 198396 |  SR142948A SR142948A | C39H51N5O6 | 685.866 | 8 / 2 | 3.6 | No |
| 200236 | 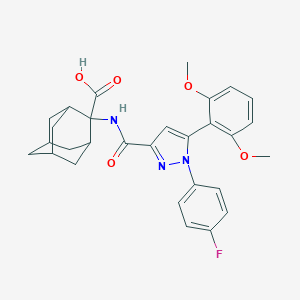 CHEMBL3290103 CHEMBL3290103 | C29H30FN3O5 | 519.573 | 7 / 2 | 5.0 | No |
| 200734 | 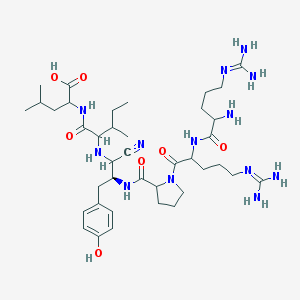 CHEMBL266981 CHEMBL266981 | C39H65N13O7 | 828.033 | 12 / 11 | -2.4 | No |
| 200735 | 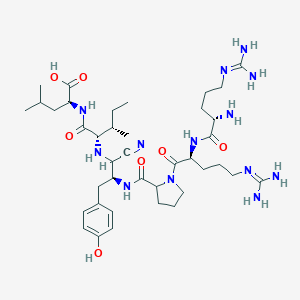 CHEMBL3143475 CHEMBL3143475 | C39H65N13O7 | 828.033 | 12 / 11 | -2.4 | No |
| 200736 | 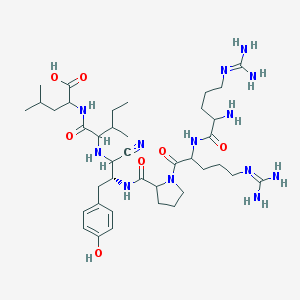 CHEMBL7384 CHEMBL7384 | C39H65N13O7 | 828.033 | 12 / 11 | -2.4 | No |
| 200737 | 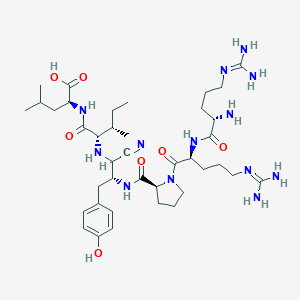 CHEMBL3143244 CHEMBL3143244 | C39H65N13O7 | 828.033 | 12 / 11 | -2.4 | No |
| 201726 |  CHEMBL2369588 CHEMBL2369588 | C41H64N6O8 | 768.997 | 9 / 7 | 2.6 | No |
| 205970 | 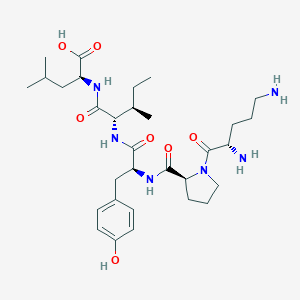 CHEMBL1790470 CHEMBL1790470 | C31H50N6O7 | 618.776 | 9 / 7 | -2.1 | No |
| 449797 | 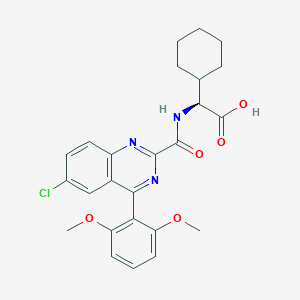 CHEMBL3356855 CHEMBL3356855 | C25H26ClN3O5 | 483.949 | 7 / 2 | 5.4 | No |
| 209279 |  CHEMBL269581 CHEMBL269581 | C37H61N7O9 | 747.935 | 12 / 8 | 1.0 | No |
| 211913 | 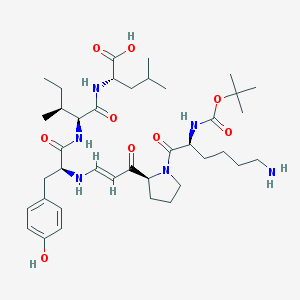 CID 44265524 CID 44265524 | C39H62N6O9 | 758.958 | 11 / 7 | 2.8 | No |
| 226799 |  CHEMBL1790457 CHEMBL1790457 | C42H66N8O9S | 859.097 | 11 / 9 | 0.7 | No |
| 229521 | 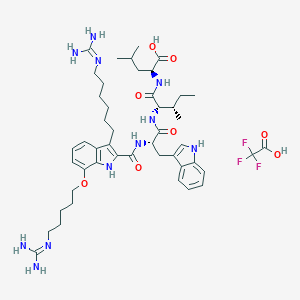 CHEMBL21892 CHEMBL21892 | C47H68F3N11O8 | 972.125 | 13 / 11 | N/A | No |
| 232006 | 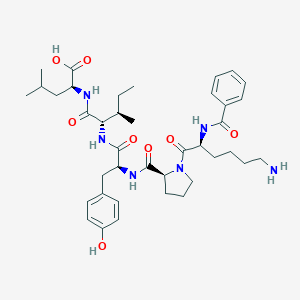 CHEMBL1790460 CHEMBL1790460 | C39H56N6O8 | 736.911 | 9 / 7 | 1.8 | No |
| 233901 | 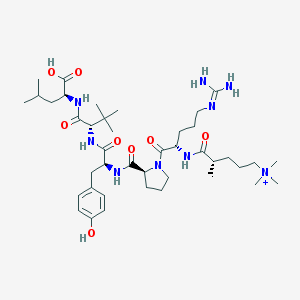 CHEMBL1172376 CHEMBL1172376 | C41H70N9O8+ | 817.066 | 9 / 8 | 2.8 | No |
| 235819 | 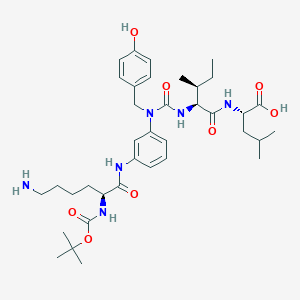 CHEMBL1794003 CHEMBL1794003 | C37H56N6O8 | 712.889 | 9 / 7 | 2.4 | No |
| 451514 |  CHEMBL3403508 CHEMBL3403508 | C24H24FN3O4 | 437.471 | 6 / 2 | 5.4 | No |
| 256348 | 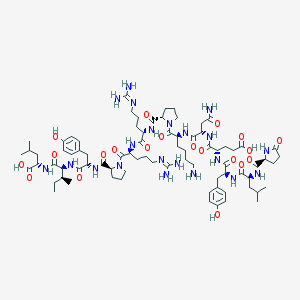 NEUROTENSIN NEUROTENSIN | C78H121N21O20 | 1672.95 | 23 / 21 | -3.7 | No |
| 256498 | 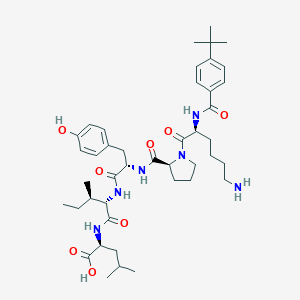 CHEMBL1790467 CHEMBL1790467 | C43H64N6O8 | 793.019 | 9 / 7 | 3.5 | No |
| 271475 | 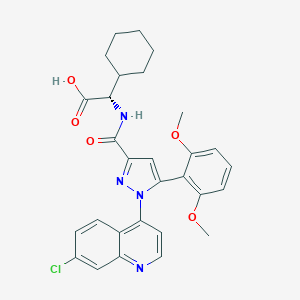 SR-48527 SR-48527 | C29H29ClN4O5 | 549.024 | 7 / 2 | 6.0 | No |
| 280822 | 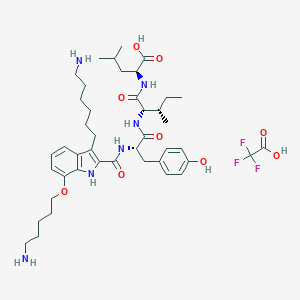 CHEMBL21521 CHEMBL21521 | C43H63F3N6O9 | 865.005 | 14 / 9 | N/A | No |
| 452938 | 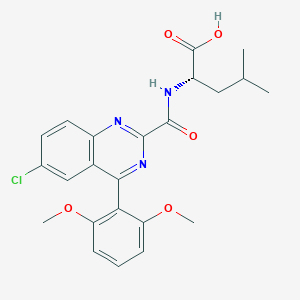 CHEMBL3356854 CHEMBL3356854 | C23H24ClN3O5 | 457.911 | 7 / 2 | 4.7 | Yes |
| 285645 | 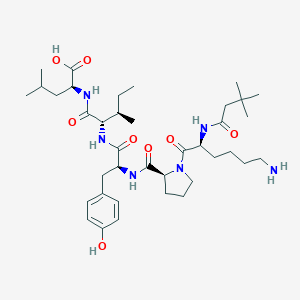 CHEMBL1790461 CHEMBL1790461 | C38H62N6O8 | 730.948 | 9 / 7 | 1.5 | No |
| 289133 | 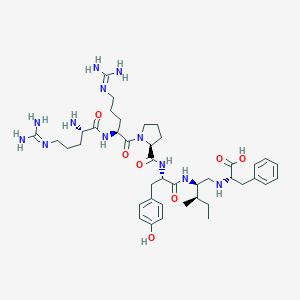 CHEMBL1793837 CHEMBL1793837 | C41H64N12O7 | 837.04 | 11 / 11 | -2.0 | No |
| 298616 | 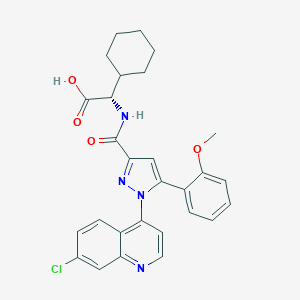 CHEMBL3290092 CHEMBL3290092 | C28H27ClN4O4 | 518.998 | 6 / 2 | 6.0 | No |
| 300712 | 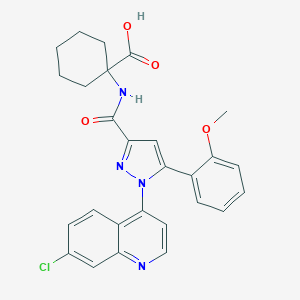 CHEMBL3290098 CHEMBL3290098 | C27H25ClN4O4 | 504.971 | 6 / 2 | 5.2 | No |
| 309941 | 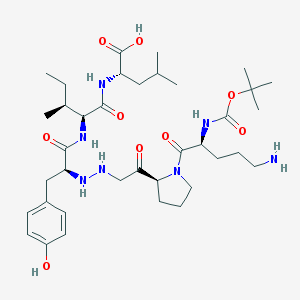 CHEMBL8899 CHEMBL8899 | C37H61N7O9 | 747.935 | 12 / 8 | 1.0 | No |
| 454393 | 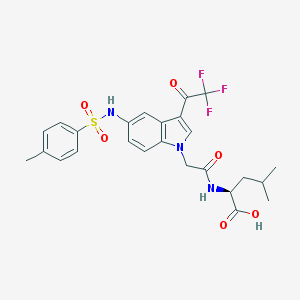 CHEMBL3326831 CHEMBL3326831 | C25H26F3N3O6S | 553.553 | 10 / 3 | 4.5 | No |
| 321296 |  CHEMBL415303 CHEMBL415303 | C42H63N11O8 | 850.035 | 11 / 10 | -2.0 | No |
| 321297 | 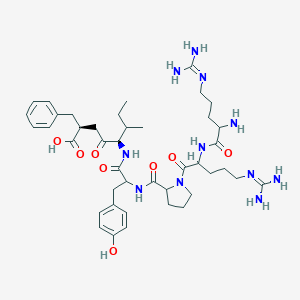 CHEMBL7405 CHEMBL7405 | C42H63N11O8 | 850.035 | 11 / 10 | -2.0 | No |
| 321298 | 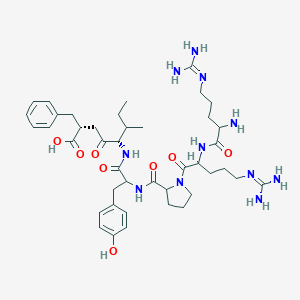 CHEMBL266131 CHEMBL266131 | C42H63N11O8 | 850.035 | 11 / 10 | -2.0 | No |
| 321299 | 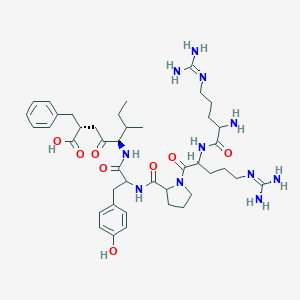 CHEMBL274093 CHEMBL274093 | C42H63N11O8 | 850.035 | 11 / 10 | -2.0 | No |
| 321300 | 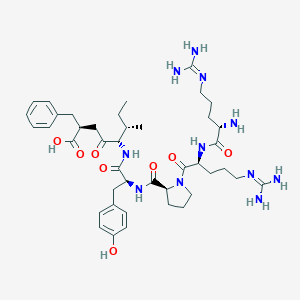 CHEMBL3143232 CHEMBL3143232 | C42H63N11O8 | 850.035 | 11 / 10 | -2.0 | No |
| 503100 | 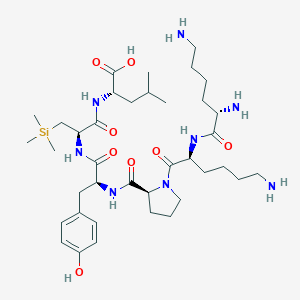 CHEMBL3622801 CHEMBL3622801 | C38H66N8O8Si | 791.079 | 11 / 9 | N/A | No |
| 554924 | 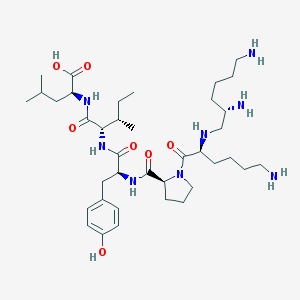 Jmv 449 Jmv 449 | C38H66N8O7 | 746.995 | 11 / 9 | -0.3 | No |
| 332476 | 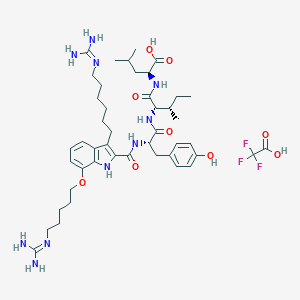 CHEMBL21448 CHEMBL21448 | C45H67F3N10O9 | 949.087 | 14 / 11 | N/A | No |
| 454913 | 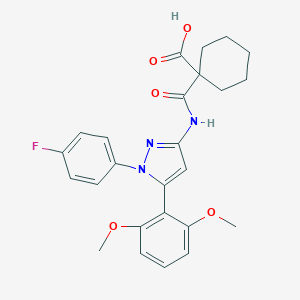 CHEMBL3403507 CHEMBL3403507 | C25H26FN3O5 | 467.497 | 7 / 2 | 5.3 | No |
| 454954 | 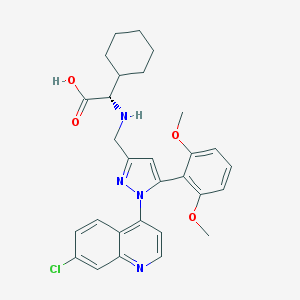 CHEMBL3403504 CHEMBL3403504 | C29H31ClN4O4 | 535.041 | 7 / 2 | 3.5 | No |
| 335307 | 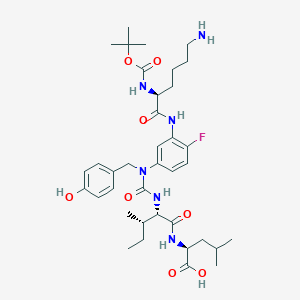 CHEMBL1793998 CHEMBL1793998 | C37H55FN6O8 | 730.879 | 10 / 7 | 2.5 | No |
| 337583 | 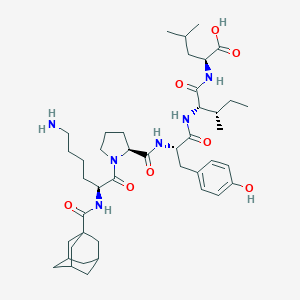 CHEMBL2372106 CHEMBL2372106 | C43H66N6O8 | 795.035 | 9 / 7 | 2.8 | No |
| 338425 | 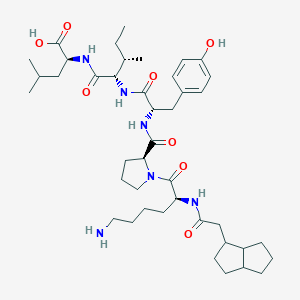 CHEMBL2369587 CHEMBL2369587 | C42H66N6O8 | 783.024 | 9 / 7 | 3.1 | No |
| 338748 | 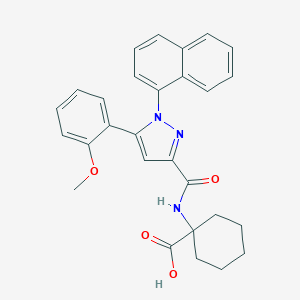 CHEMBL3290102 CHEMBL3290102 | C28H27N3O4 | 469.541 | 5 / 2 | 5.6 | No |
| 363632 | 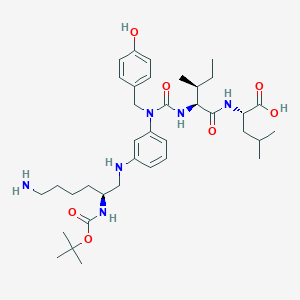 CHEMBL1793833 CHEMBL1793833 | C37H58N6O7 | 698.906 | 9 / 7 | 3.2 | No |
| 377949 | 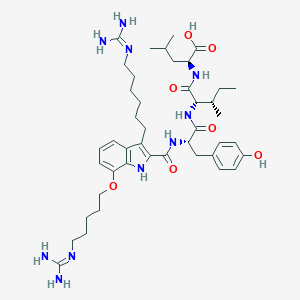 CHEMBL21448 CHEMBL21448 | C43H66N10O7 | 835.064 | 9 / 10 | 5.0 | No |
| 385048 | 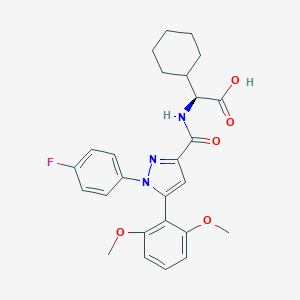 CHEMBL3290104 CHEMBL3290104 | C26H28FN3O5 | 481.524 | 7 / 2 | 5.2 | No |
| 456985 | 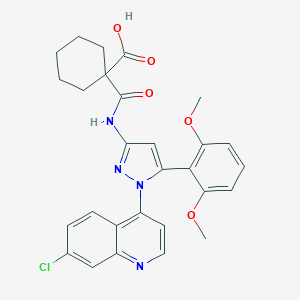 CHEMBL3403505 CHEMBL3403505 | C28H27ClN4O5 | 534.997 | 7 / 2 | 6.1 | No |
| 386453 | 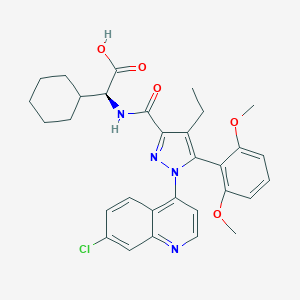 CHEMBL3290094 CHEMBL3290094 | C31H33ClN4O5 | 577.078 | 7 / 2 | 6.8 | No |
| 392930 | 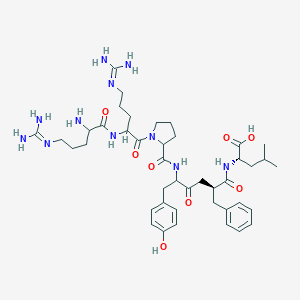 CHEMBL7674 CHEMBL7674 | C42H63N11O8 | 850.035 | 11 / 10 | -3.5 | No |
| 392931 | 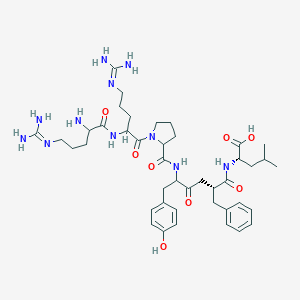 CHEMBL269292 CHEMBL269292 | C42H63N11O8 | 850.035 | 11 / 10 | -3.5 | No |
| 392932 | 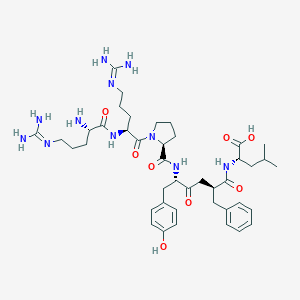 CHEMBL3143334 CHEMBL3143334 | C42H63N11O8 | 850.035 | 11 / 10 | -3.5 | No |
| 393642 | 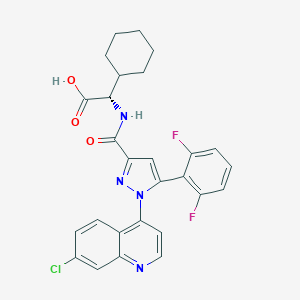 CHEMBL3290093 CHEMBL3290093 | C27H23ClF2N4O3 | 524.953 | 7 / 2 | 6.3 | No |
| 401277 | 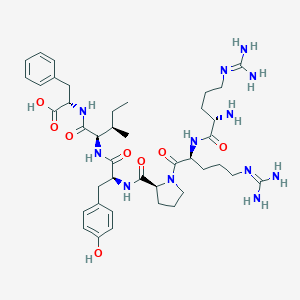 CHEMBL1793839 CHEMBL1793839 | C41H62N12O8 | 851.023 | 11 / 11 | -2.2 | No |
| 457734 |  CHEMBL3326832 CHEMBL3326832 | C23H27N3O5S | 457.545 | 6 / 3 | 3.7 | Yes |
| 405655 | 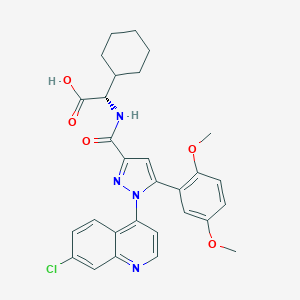 CHEMBL3290090 CHEMBL3290090 | C29H29ClN4O5 | 549.024 | 7 / 2 | 6.0 | No |
| 459228 |  CHEMBL3403506 CHEMBL3403506 | C29H29N3O5 | 499.567 | 6 / 2 | 6.5 | No |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218