

You can:
| Name | Taste receptor type 1 member 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | TAS1R2 |
| Synonym | taste receptor, type 1, member 2 taste receptor TAS1R2 T1R2 Sweet taste receptor T1R2 [ Show all ] |
| Disease | N/A |
| Length | 839 |
| Amino acid sequence | MGPRAKTISSLFFLLWVLAEPAENSDFYLPGDYLLGGLFSLHANMKGIVHLNFLQVPMCKEYEVKVIGYNLMQAMRFAVEEINNDSSLLPGVLLGYEIVDVCYISNNVQPVLYFLAHEDNLLPIQEDYSNYISRVVAVIGPDNSESVMTVANFLSLFLLPQITYSAISDELRDKVRFPALLRTTPSADHHIEAMVQLMLHFRWNWIIVLVSSDTYGRDNGQLLGERVARRDICIAFQETLPTLQPNQNMTSEERQRLVTIVDKLQQSTARVVVVFSPDLTLYHFFNEVLRQNFTGAVWIASESWAIDPVLHNLTELRHLGTFLGITIQSVPIPGFSEFREWGPQAGPPPLSRTSQSYTCNQECDNCLNATLSFNTILRLSGERVVYSVYSAVYAVAHALHSLLGCDKSTCTKRVVYPWQLLEEIWKVNFTLLDHQIFFDPQGDVALHLEIVQWQWDRSQNPFQSVASYYPLQRQLKNIQDISWHTINNTIPMSMCSKRCQSGQKKKPVGIHVCCFECIDCLPGTFLNHTEDEYECQACPNNEWSYQSETSCFKRQLVFLEWHEAPTIAVALLAALGFLSTLAILVIFWRHFQTPIVRSAGGPMCFLMLTLLLVAYMVVPVYVGPPKVSTCLCRQALFPLCFTICISCIAVRSFQIVCAFKMASRFPRAYSYWVRYQGPYVSMAFITVLKMVIVVIGMLATGLSPTTRTDPDDPKITIVSCNPNYRNSLLFNTSLDLLLSVVGFSFAYMGKELPTNYNEAKFITLSMTFYFTSSVSLCTFMSAYSGVLVTIVDLLVTVLNLLAISLGYFGPKCYMILFYPERNTPAYFNSMIQGYTMRRD |
| UniProt | Q8TE23 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | N/A |
| IUPHAR | N/A |
| DrugBank | BE0000395 |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 548513 | 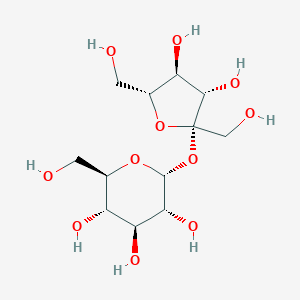 sucrose sucrose | C12H22O11 | 342.297 | 11 / 8 | -3.7 | No |
| 128619 | 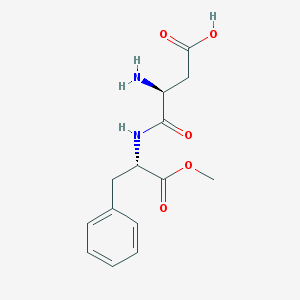 aspartame aspartame | C14H18N2O5 | 294.307 | 6 / 3 | -2.7 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218