
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | |
| | | | | | | | | | | | | | | | | | | | | |
| MAATASPQPLATEDADSENSSFYYYDYLDEVAFMLCRKDAVVSFGKVFLPVFYSLIFVLGLSGNLLLLMVLLRYVPRRRMVEIYLLNLAISNLLFLVTLPFWGISVAWHWVFGSFLCKMVSTLYTINFYSGIFFISCMSLDKYLEIVHAQPYHRLRTRAKSLLLATIVWAVSLAVSIPDMVFVQTHENPKGVWNCHADFGGHGTIWKLFLRFQQNLLGFLLPLLAMIFFYSRIGCVLVRLRPAGQGRALKIAAALVVAFFVLWFPYNLTLFLHTLLDLQVFGNCEVSQHLDYALQVTESIAFLHCCFSPILYAFSSHRFRQYLKAFLAAVLGWHLAPGTAQASLSSCSESSILTAQEEMTGMNDLGERQSENYPNKEDVGNKSA | |
| CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCCSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC | |
| 999889987778888877777777677777655678752036489999999999999999998999899877346768699999999999999999989999999758997966789999999999999999999999999678987102134465630364202999999999999999985217889994798334896207899999999999999999999999999999999634886676179999999999999858999999999999856789846999999999999999999998878988814888999999998876367778888888888899998877777777888867788888999998777789 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | |
| | | | | | | | | | | | | | | | | | | | | |
| MAATASPQPLATEDADSENSSFYYYDYLDEVAFMLCRKDAVVSFGKVFLPVFYSLIFVLGLSGNLLLLMVLLRYVPRRRMVEIYLLNLAISNLLFLVTLPFWGISVAWHWVFGSFLCKMVSTLYTINFYSGIFFISCMSLDKYLEIVHAQPYHRLRTRAKSLLLATIVWAVSLAVSIPDMVFVQTHENPKGVWNCHADFGGHGTIWKLFLRFQQNLLGFLLPLLAMIFFYSRIGCVLVRLRPAGQGRALKIAAALVVAFFVLWFPYNLTLFLHTLLDLQVFGNCEVSQHLDYALQVTESIAFLHCCFSPILYAFSSHRFRQYLKAFLAAVLGWHLAPGTAQASLSSCSESSILTAQEEMTGMNDLGERQSENYPNKEDVGNKSA | |
| 774443443333543433332232331233443320436314300100002101200230232120001000112331100000000001010000000000000002001102000120000122111000000000030000000000202322322101000000110000001000000202437622110103035642201000111111102331220012000000110131464633000000000000000012210100001002313224414123001100000100002000310000000044014101300340043324564444445444543444445533435534535655436555346648 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 380 | | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCCSSSSCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCC MAATASPQPLATEDADSENSSFYYYDYLDEVAFMLCRKDAVVSFGKVFLPVFYSLIFVLGLSGNLLLLMVLLRYVPRRRMVEIYLLNLAISNLLFLVTLPFWGISVAWHWVFGSFLCKMVSTLYTINFYSGIFFISCMSLDKYLEIVHAQPYHRLRTRAKSLLLATIVWAVSLAVSIPDMVFVQTHENPKGVWNCHADFGGHGTIWKLFLRFQQNLLGFLLPLLAMIFFYSRIGCVLVRLRPAGQGRALKIAAALVVAFFVLWFPYNLTLFLHTLLDLQVFGNCEVSQHLDYALQVTESIAFLHCCFSPILYAFSSHRFRQYLKAFLAAVLGWHLAPGTAQASLSSCSESSILTAQEEMTGMNDLGERQSENYPNKEDVGNKSA | |||||||||||||||||||||||||
| 1 | 4mbsA | 0.38 | 0.29 | 0.76 | 3.35 | Download | ----------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEG-LHYTCSSHFPYSQQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ-------------------------------------------------------- | |||||||||||||||||||
| 2 | 4mbsA | 0.38 | 0.30 | 0.76 | 4.03 | Download | ----------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQK-EGLHYTCSSHFPSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMEEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFF--------------------------------------------------------- | |||||||||||||||||||
| 3 | 4mbsA | 0.38 | 0.29 | 0.76 | 3.70 | Download | ----------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEGL-HYTCSSHFPYQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEEEKKRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ-------------------------------------------------------- | |||||||||||||||||||
| 4 | 2ziy | 0.19 | 0.19 | 0.90 | 1.55 | Download | -------------DLR-DNETW-WYNPSIIVHPHWREFDQVPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVFPLMTISCLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGWGAYTLEGVLCNCSFDYISRDSTTRSN-ILCMFILGFFGPILIIFFCYFNIVMSVSNHEKNAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLE--------WVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQFDDKETE--DD--KDAE--T---EIPAGESSDAAPSADAAQ----MKE- | |||||||||||||||||||
| 5 | 5t1a | 0.39 | 0.30 | 0.73 | 1.19 | Download | ----------------------------------------VKQIGAQLLPPLYSLVFIFGFVGNMLVVLILINCKKLKCLTDIYLLNLAISDLLFLITLPLWAHSAANEWVFGNAMCKLFTGLYHIGYFGGIFFIILLTIDRYLAIVHAVFALKARTVTFGVVTSVITWLVAVFASVPGIIFTKQKE--DSVYVCGPYFPR---GWNNFHTIMRNILGLVLPLLIMVICYSGISRASKSRIPSREKKAVRVIFTIMIVYFLFWTPYNIVILLNTFQEFFGLSNCESTSQLDQATQVTETLGMTHCCINPIIYAFVGEKFRRYLSVFF--------------------------------------------------------- | |||||||||||||||||||
| 6 | 4mbsA | 0.38 | 0.29 | 0.76 | 3.30 | Download | -----------------------------------CQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEGL-HYTCSSHFPSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ-------------------------------------------------------- | |||||||||||||||||||
| 7 | 5t1a | 0.39 | 0.30 | 0.73 | 1.72 | Download | ------------------------------------------QIGAQLLPPLYSLVFIFGFVGNMLVVLILINCKKLKCLTDIYLLNLAISDLLFLITLPLWAHSAANEWVFGNAMCKLFTGLYHIGYFGGIFFIILLTIDRYLAIVHAVFALKARTVTFGVVTSVITWLVAVFASVPGIIFTKQKE--DSVYVCGPYFPRG---WNNFHTIMRNILGLVLPLLIMVICYSGISRASKSRPPSREKKAVRVIFTIMIVYFLFWTPYNIVILLNTFQEFFGLSNCESTSQLDQATQVTETLGMTHCCINPIIYAFVGEKFRRYLSVFF--------------------------------------------------------- | |||||||||||||||||||
| 8 | 4mbsA | 0.37 | 0.29 | 0.76 | 4.94 | Download | ----------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEGLHYTCSSHFPYSQYQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLREK--KRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ-------------------------------------------------------- | |||||||||||||||||||
| 9 | 4mbsA | 0.38 | 0.29 | 0.76 | 3.64 | Download | ----------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAAAQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKE-GLHYTCSSHFPYSQQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLRMKEEEKHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ-------------------------------------------------------- | |||||||||||||||||||
| 10 | 5c1mA | 0.24 | 0.21 | 0.74 | 3.97 | Download | ------------------------------GSHSLPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVRYTKMKTATNIYIFNLALADALATSTLPFQSVNYLMGWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKY-RQGSIDCTLTFSHPTWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALI------TIPETTFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCF------------------------------------------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
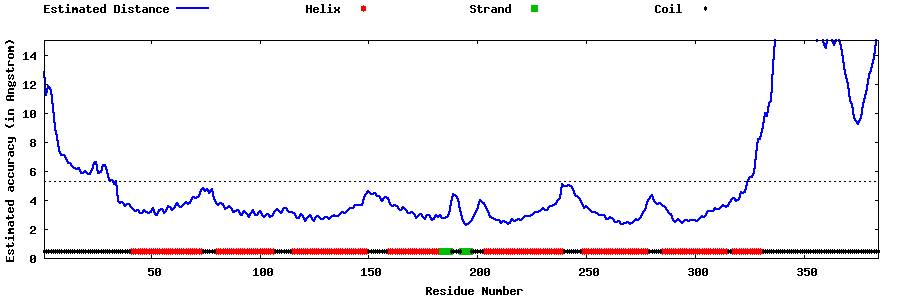
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||