

You can:
| Name | CHEMBL163813 |
|---|---|
| Molecular formula | C26H34ClNO3S |
| IUPAC name | propyl 2-(3-chlorophenyl)-4,6-diethyl-5-hexylsulfanylcarbonylpyridine-3-carboxylate |
| Molecular weight | 476.072 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 0 |
| XlogP | 8.2 |
| Synonyms | BDBM50074227 2-(3-Chloro-phenyl)-4,6-diethyl-5-hexylsulfanylcarbonyl-nicotinic acid propyl ester |
| Inchi Key | APMWCJWGZQDHJU-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C26H34ClNO3S/c1-5-9-10-11-16-32-26(30)22-20(7-3)23(25(29)31-15-6-2)24(28-21(22)8-4)18-13-12-14-19(27)17-18/h12-14,17H,5-11,15-16H2,1-4H3 |
| PubChem CID | 10719352 |
| ChEMBL | CHEMBL163813 |
| IUPHAR | N/A |
| BindingDB | 50074227 |
| DrugBank | N/A |
Structure | 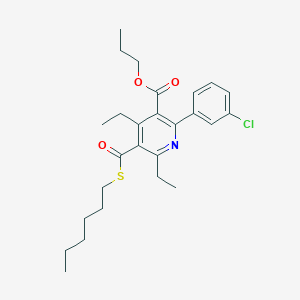 |
| Lipinski's druglikeness | This ligand has a partition coefficient log P greater than 5. |
You can:
| GLASS ID | Name | UniProt | Gene | Species | Length |
|---|---|---|---|---|---|
| 11135 | Adenosine receptor A1 | P25099 | Adora1 | Rattus norvegicus (Rat) | 326 |
| 11136 | Adenosine receptor A2a | P30543 | Adora2a | Rattus norvegicus (Rat) | 410 |
| 11134 | Adenosine receptor A3 | P0DMS8 | ADORA3 | Homo sapiens (Human) | 318 |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218