

You can:
| Name | G-protein coupled bile acid receptor 1 |
|---|---|
| Species | Mus musculus (Mouse) |
| Gene | Gpbar1 |
| Synonym | membrane-type receptor for bile acids M-BAR hGPCR19 GPR131 GPCR19 [ Show all ] |
| Disease | N/A for non-human GPCRs |
| Length | 329 |
| Amino acid sequence | MMTPNSTELSAIPMGVLGLSLALASLIVIANLLLALGIALDRHLRSPPAGCFFLSLLLAGLLTGLALPMLPGLWSRNHQGYWSCLLLHLTPNFCFLSLLANLLLVHGERYMAVLQPLRPHGSVRLALFLTWVSSLFFASLPALGWNHWSPDANCSSQAVFPAPYLYLEVYGLLLPAVGATALLSVRVLATAHRQLCEIRRLERAVCRDVPSTLARALTWRQARAQAGATLLFLLCWGPYVATLLLSVLAYERRPPLGPGTLLSLISLGSTSAAAVPVAMGLGDQRYTAPWRTAAQRCLRVLRGRAKRDNPGPSTAYHTSSQCSIDLDLN |
| UniProt | Q80SS6 |
| Protein Data Bank | N/A |
| GPCR-HGmod model | N/A |
| 3D structure model | No available structures or models |
| BioLiP | N/A |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL1255150 |
| IUPHAR | 37 |
| DrugBank | N/A |
You can:
| GLASS ID | Molecule | Formula | Molecular weight | H-bond acceptor / donor | XlogP | Lipinski's druglikeness |
|---|---|---|---|---|---|---|
| 9742 | 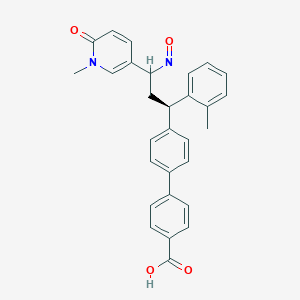 BDBM50437875 BDBM50437875 | C29H26N2O4 | 466.537 | 5 / 1 | 4.6 | Yes |
| 9743 |  BDBM50437874 BDBM50437874 | C29H26N2O4 | 466.537 | 5 / 1 | 4.6 | Yes |
| 557580 |  CHEMBL1257491 CHEMBL1257491 | C22H19BrN2OS | 439.371 | 4 / 2 | 5.2 | No |
| 12528 | 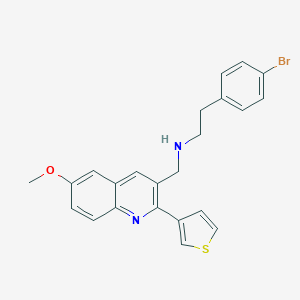 CHEMBL1257611 CHEMBL1257611 | C23H21BrN2OS | 453.398 | 4 / 1 | 5.5 | No |
| 13427 | 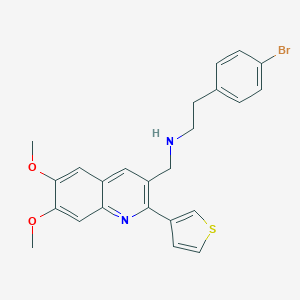 CHEMBL1257733 CHEMBL1257733 | C24H23BrN2O2S | 483.424 | 5 / 1 | 5.5 | No |
| 19911 | 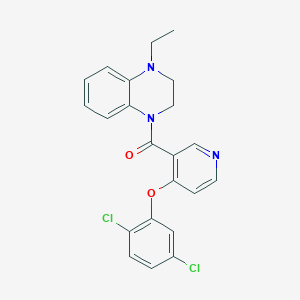 CHEMBL2181235 CHEMBL2181235 | C22H19Cl2N3O2 | 428.313 | 4 / 0 | 4.9 | Yes |
| 20329 | 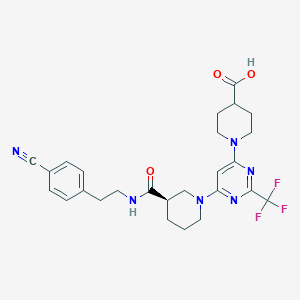 CHEMBL3234580 CHEMBL3234580 | C26H29F3N6O3 | 530.552 | 11 / 2 | 3.5 | No |
| 23964 |  cholic acid cholic acid | C24H40O5 | 408.579 | 5 / 4 | 3.6 | Yes |
| 24739 | 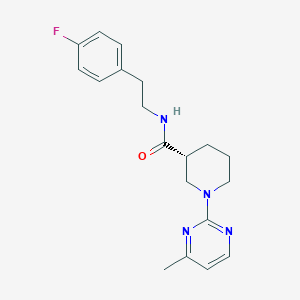 CHEMBL3234551 CHEMBL3234551 | C19H23FN4O | 342.418 | 5 / 1 | 2.9 | Yes |
| 31655 | 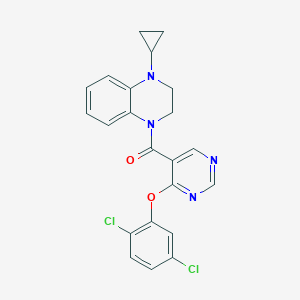 CHEMBL2181256 CHEMBL2181256 | C22H18Cl2N4O2 | 441.312 | 5 / 0 | 4.8 | Yes |
| 35137 | 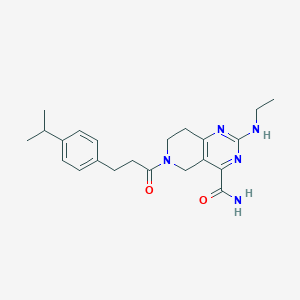 CHEMBL2331646 CHEMBL2331646 | C22H29N5O2 | 395.507 | 5 / 2 | 2.5 | Yes |
| 35984 | 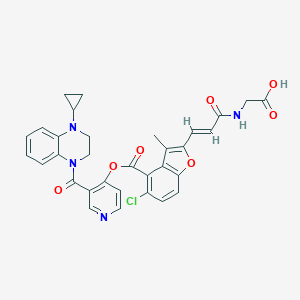 CHEMBL3290734 CHEMBL3290734 | C32H27ClN4O7 | 615.039 | 9 / 2 | 4.5 | No |
| 37471 | 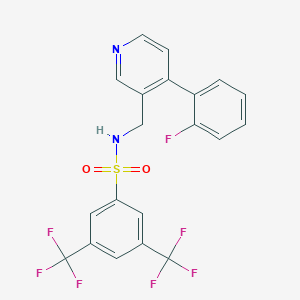 CHEMBL2435927 CHEMBL2435927 | C20H13F7N2O2S | 478.385 | 11 / 1 | 4.8 | No |
| 558519 | 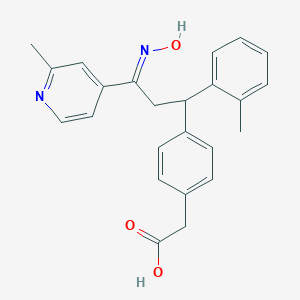 SCHEMBL10240963 SCHEMBL10240963 | C24H24N2O3 | 388.467 | 5 / 2 | 4.4 | Yes |
| 46478 | 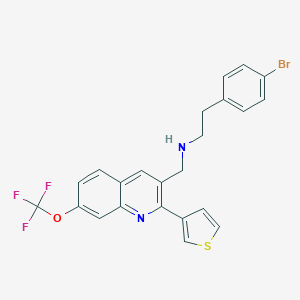 CHEMBL1257380 CHEMBL1257380 | C23H18BrF3N2OS | 507.369 | 7 / 1 | 6.7 | No |
| 443925 |  CHEMBL3421893 CHEMBL3421893 | C28H27Cl2N3O4 | 540.441 | 6 / 0 | 5.6 | No |
| 57101 | 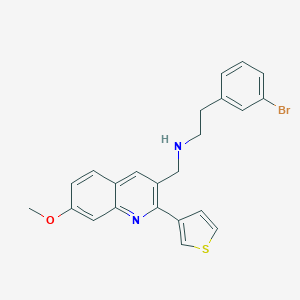 CHEMBL1258424 CHEMBL1258424 | C23H21BrN2OS | 453.398 | 4 / 1 | 5.5 | No |
| 58669 | 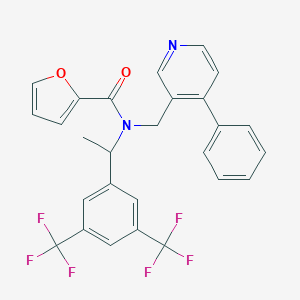 CHEMBL2435945 CHEMBL2435945 | C27H20F6N2O2 | 518.459 | 9 / 0 | 6.5 | No |
| 58697 | 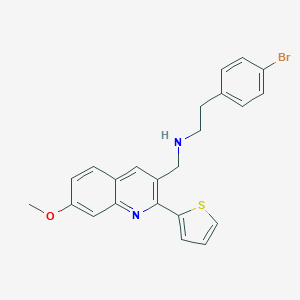 CHEMBL1258530 CHEMBL1258530 | C23H21BrN2OS | 453.398 | 4 / 1 | 5.6 | No |
| 444238 |  CHEMBL3421894 CHEMBL3421894 | C26H23Cl2N3O4 | 512.387 | 6 / 1 | 4.9 | No |
| 444397 | 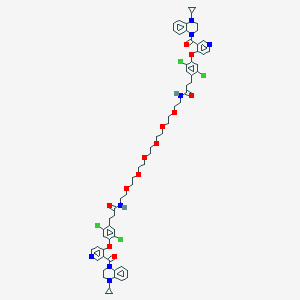 CHEMBL3421903 CHEMBL3421903 | C66H74Cl4N8O12 | 1313.16 | 16 / 2 | 8.1 | No |
| 69057 | 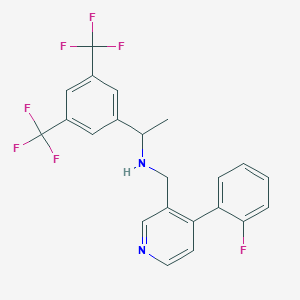 CHEMBL2435921 CHEMBL2435921 | C22H17F7N2 | 442.381 | 9 / 1 | 5.6 | No |
| 69509 | 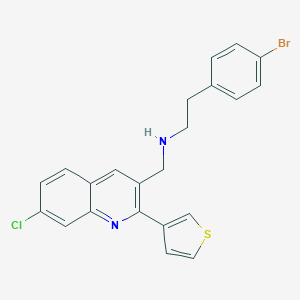 CHEMBL1257259 CHEMBL1257259 | C22H18BrClN2S | 457.814 | 3 / 1 | 6.2 | No |
| 70106 | 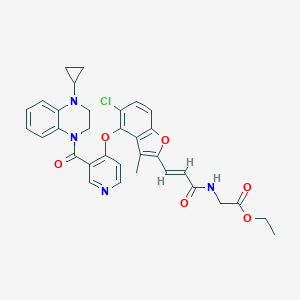 CHEMBL3290733 CHEMBL3290733 | C33H31ClN4O6 | 615.083 | 8 / 1 | 5.3 | No |
| 75413 | 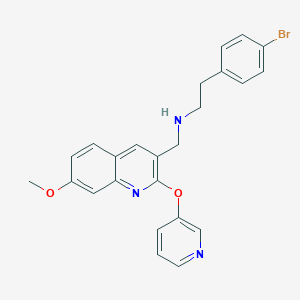 CHEMBL1258877 CHEMBL1258877 | C24H22BrN3O2 | 464.363 | 5 / 1 | 5.0 | Yes |
| 444871 | 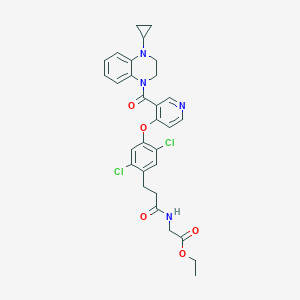 CHEMBL3421896 CHEMBL3421896 | C30H30Cl2N4O5 | 597.493 | 7 / 1 | 4.9 | No |
| 80798 | 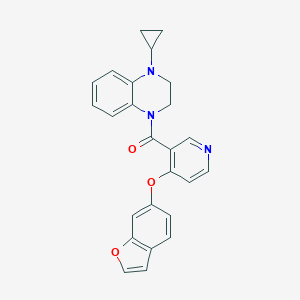 CHEMBL3290707 CHEMBL3290707 | C25H21N3O3 | 411.461 | 5 / 0 | 4.3 | Yes |
| 553673 | 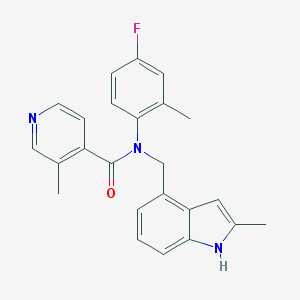 GTPL9078 GTPL9078 | C24H22FN3O | 387.458 | 3 / 1 | 4.7 | Yes |
| 560092 | 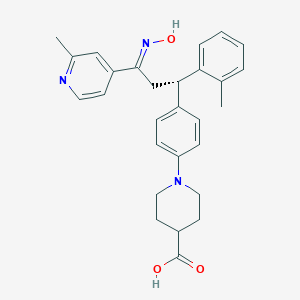 SCHEMBL1389953 SCHEMBL1389953 | C28H31N3O3 | 457.574 | 6 / 2 | 5.0 | Yes |
| 560094 |  SCHEMBL364189 SCHEMBL364189 | C28H31N3O3 | 457.574 | 6 / 2 | 5.0 | Yes |
| 91795 | 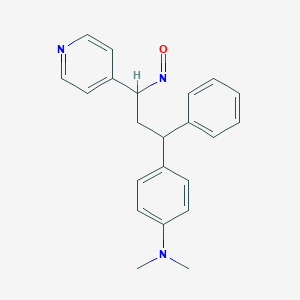 BDBM50437890 BDBM50437890 | C22H23N3O | 345.446 | 4 / 0 | 4.1 | Yes |
| 560164 | 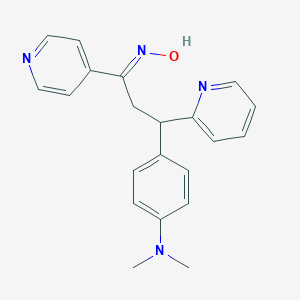 SCHEMBL364486 SCHEMBL364486 | C21H22N4O | 346.434 | 5 / 1 | 3.2 | Yes |
| 94501 | 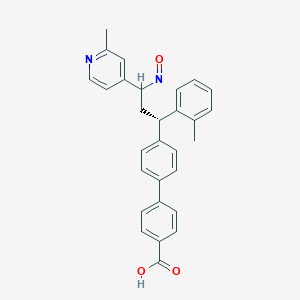 BDBM50437878 BDBM50437878 | C29H26N2O3 | 450.538 | 5 / 1 | 5.9 | No |
| 94503 | 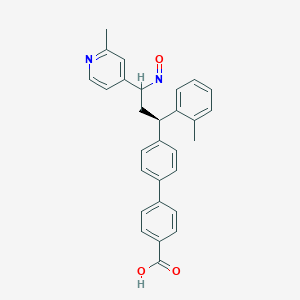 BDBM50437879 BDBM50437879 | C29H26N2O3 | 450.538 | 5 / 1 | 5.9 | No |
| 560255 |  SCHEMBL1390210 SCHEMBL1390210 | C21H19BrN2O | 395.3 | 3 / 1 | 5.2 | No |
| 95984 | 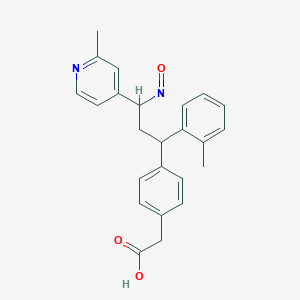 BDBM50437881 BDBM50437881 | C24H24N2O3 | 388.467 | 5 / 1 | 4.2 | Yes |
| 97272 | 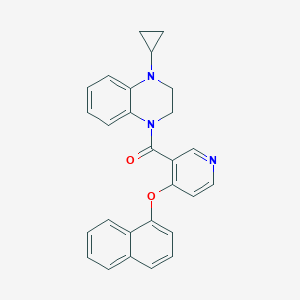 CHEMBL3290700 CHEMBL3290700 | C27H23N3O2 | 421.5 | 4 / 0 | 5.1 | No |
| 99082 | 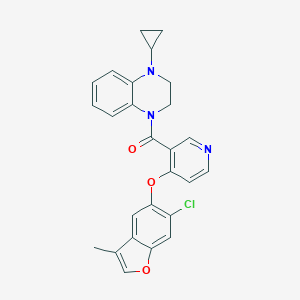 CHEMBL3290717 CHEMBL3290717 | C26H22ClN3O3 | 459.93 | 5 / 0 | 5.2 | No |
| 100343 |  CHEMBL2181223 CHEMBL2181223 | C24H23N3O3 | 401.466 | 5 / 0 | 3.8 | Yes |
| 100374 | 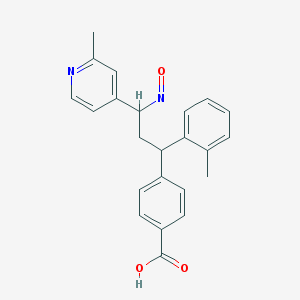 BDBM50437882 BDBM50437882 | C23H22N2O3 | 374.44 | 5 / 1 | 4.3 | Yes |
| 103555 | 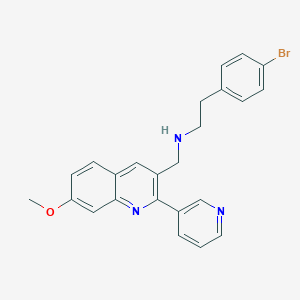 CHEMBL1258759 CHEMBL1258759 | C24H22BrN3O | 448.364 | 4 / 1 | 4.8 | Yes |
| 106559 | 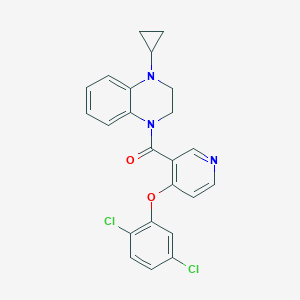 CHEMBL2181239 CHEMBL2181239 | C23H19Cl2N3O2 | 440.324 | 4 / 0 | 5.1 | No |
| 560594 | 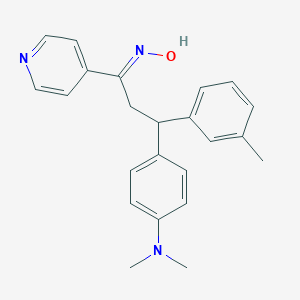 SCHEMBL10240836 SCHEMBL10240836 | C23H25N3O | 359.473 | 4 / 1 | 4.6 | Yes |
| 560688 | 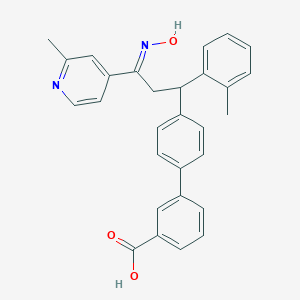 SCHEMBL10240962 SCHEMBL10240962 | C29H26N2O3 | 450.538 | 5 / 2 | 6.1 | No |
| 113325 | 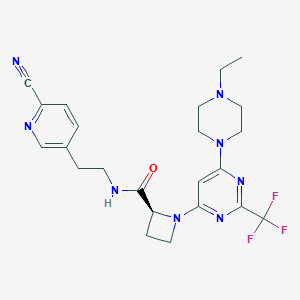 CHEMBL3234579 CHEMBL3234579 | C23H27F3N8O | 488.519 | 11 / 1 | 2.7 | No |
| 116580 | 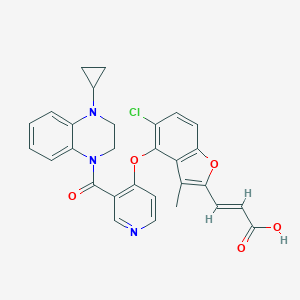 CHEMBL3290732 CHEMBL3290732 | C29H24ClN3O5 | 529.977 | 7 / 1 | 5.2 | No |
| 119589 | 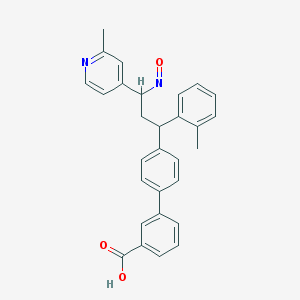 BDBM50437880 BDBM50437880 | C29H26N2O3 | 450.538 | 5 / 1 | 5.9 | No |
| 122817 | 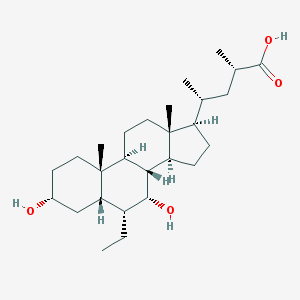 D0MZ9Q D0MZ9Q | C27H46O4 | 434.661 | 4 / 3 | 6.3 | No |
| 129218 |  CHEMBL1258423 CHEMBL1258423 | C23H21BrN2OS | 453.398 | 4 / 1 | 5.5 | No |
| 130816 | 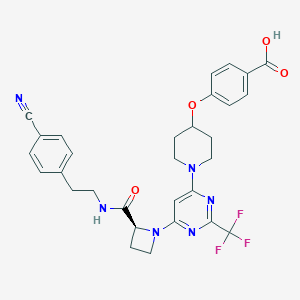 CHEMBL3234868 CHEMBL3234868 | C30H29F3N6O4 | 594.595 | 12 / 2 | 5.0 | No |
| 446982 | 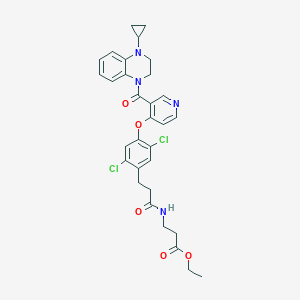 CHEMBL3421897 CHEMBL3421897 | C31H32Cl2N4O5 | 611.52 | 7 / 1 | 4.8 | No |
| 136538 | 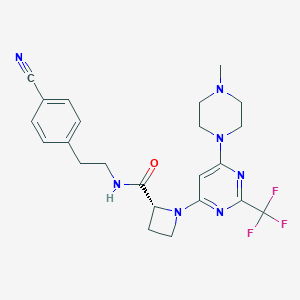 CHEMBL3234578 CHEMBL3234578 | C23H26F3N7O | 473.504 | 10 / 1 | 3.0 | Yes |
| 136540 | 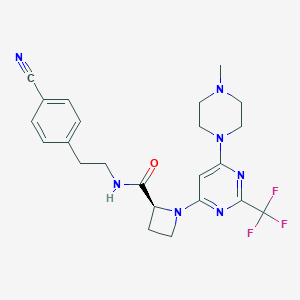 CHEMBL3234577 CHEMBL3234577 | C23H26F3N7O | 473.504 | 10 / 1 | 3.0 | Yes |
| 137295 | 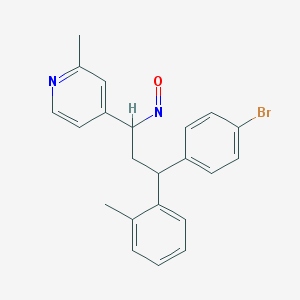 BDBM50437884 BDBM50437884 | C22H21BrN2O | 409.327 | 3 / 0 | 5.4 | No |
| 138663 | 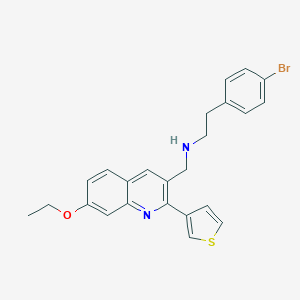 CHEMBL1257260 CHEMBL1257260 | C24H23BrN2OS | 467.425 | 4 / 1 | 5.9 | No |
| 140900 | 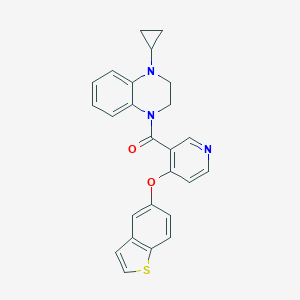 CHEMBL3290711 CHEMBL3290711 | C25H21N3O2S | 427.522 | 5 / 0 | 4.9 | Yes |
| 142667 | 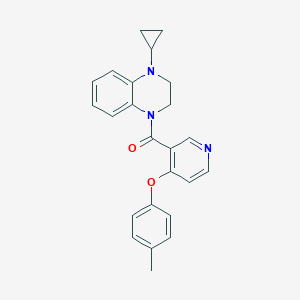 CHEMBL2181225 CHEMBL2181225 | C24H23N3O2 | 385.467 | 4 / 0 | 4.2 | Yes |
| 147594 | 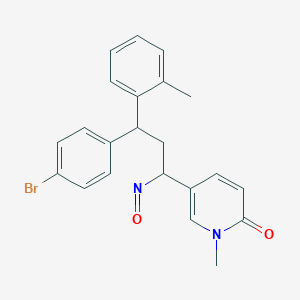 BDBM50437883 BDBM50437883 | C22H21BrN2O2 | 425.326 | 3 / 0 | 4.1 | Yes |
| 151172 | 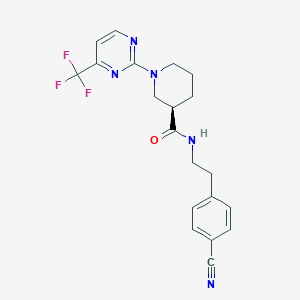 CHEMBL3234562 CHEMBL3234562 | C20H20F3N5O | 403.409 | 8 / 1 | 3.0 | Yes |
| 151245 | 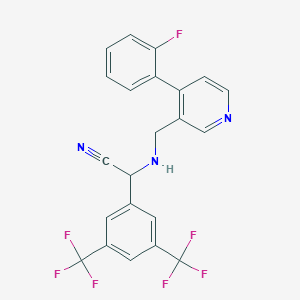 CHEMBL2435924 CHEMBL2435924 | C22H14F7N3 | 453.364 | 10 / 1 | 5.2 | No |
| 153596 | 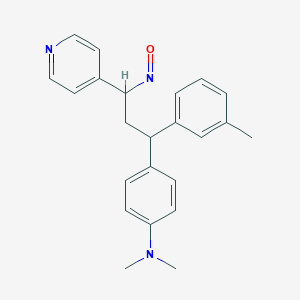 BDBM50437888 BDBM50437888 | C23H25N3O | 359.473 | 4 / 0 | 4.4 | Yes |
| 155531 | 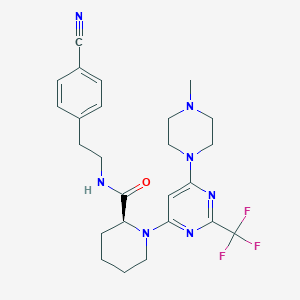 CHEMBL3234574 CHEMBL3234574 | C25H30F3N7O | 501.558 | 10 / 1 | 3.7 | No |
| 157629 | 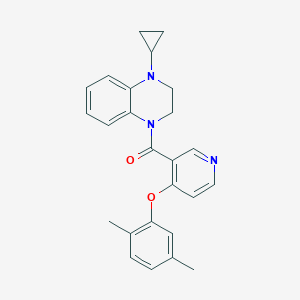 CHEMBL2181226 CHEMBL2181226 | C25H25N3O2 | 399.494 | 4 / 0 | 4.6 | Yes |
| 160308 | 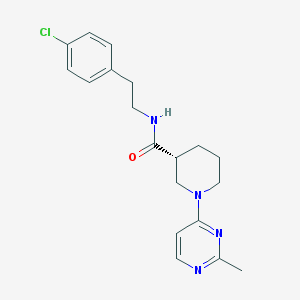 CHEMBL3234558 CHEMBL3234558 | C19H23ClN4O | 358.87 | 4 / 1 | 3.4 | Yes |
| 160655 | 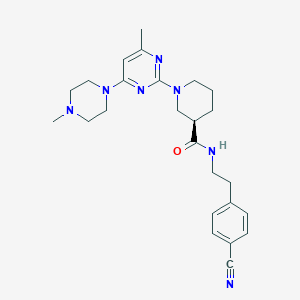 CHEMBL3234564 CHEMBL3234564 | C25H33N7O | 447.587 | 7 / 1 | 2.8 | Yes |
| 164977 | 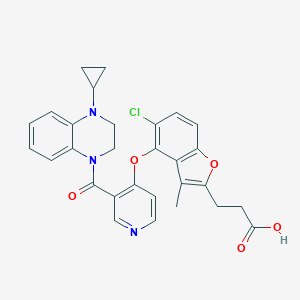 CHEMBL3290736 CHEMBL3290736 | C29H26ClN3O5 | 531.993 | 7 / 1 | 5.0 | No |
| 166659 | 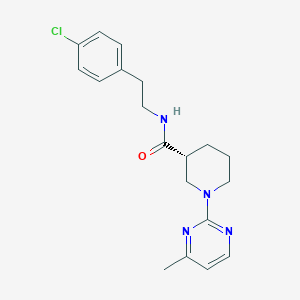 CHEMBL3234549 CHEMBL3234549 | C19H23ClN4O | 358.87 | 4 / 1 | 3.4 | Yes |
| 168116 | 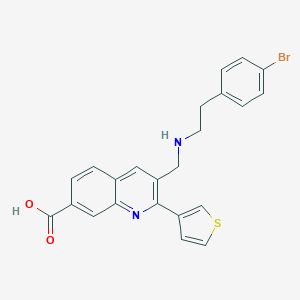 CHEMBL1257381 CHEMBL1257381 | C23H19BrN2O2S | 467.381 | 5 / 2 | 2.8 | Yes |
| 168824 | 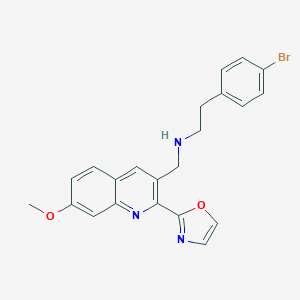 CHEMBL1257142 CHEMBL1257142 | C22H20BrN3O2 | 438.325 | 5 / 1 | 4.3 | Yes |
| 169820 | 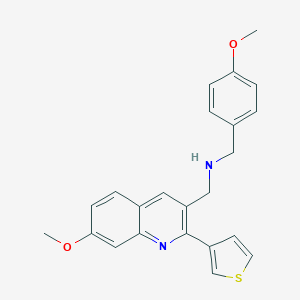 CHEMBL1258196 CHEMBL1258196 | C23H22N2O2S | 390.501 | 5 / 1 | 4.3 | Yes |
| 171258 | 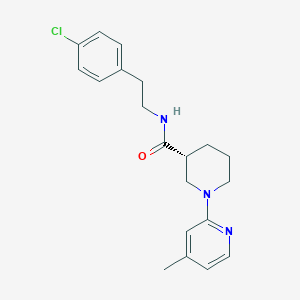 CHEMBL3234560 CHEMBL3234560 | C20H24ClN3O | 357.882 | 3 / 1 | 4.0 | Yes |
| 175269 | 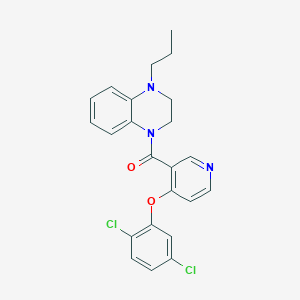 CHEMBL2181236 CHEMBL2181236 | C23H21Cl2N3O2 | 442.34 | 4 / 0 | 5.4 | No |
| 175529 | 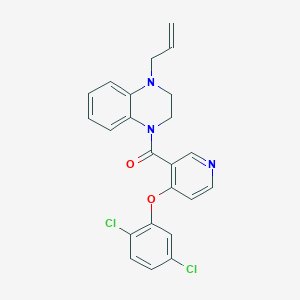 CHEMBL2181238 CHEMBL2181238 | C23H19Cl2N3O2 | 440.324 | 4 / 0 | 5.2 | No |
| 178776 | 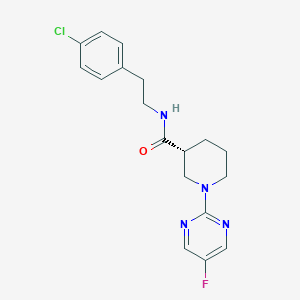 CHEMBL3234554 CHEMBL3234554 | C18H20ClFN4O | 362.833 | 5 / 1 | 3.1 | Yes |
| 180952 | 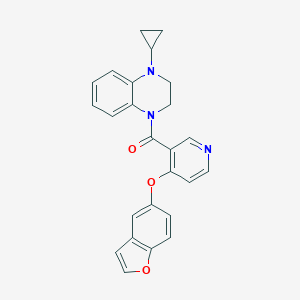 CHEMBL3290706 CHEMBL3290706 | C25H21N3O3 | 411.461 | 5 / 0 | 4.3 | Yes |
| 181032 | 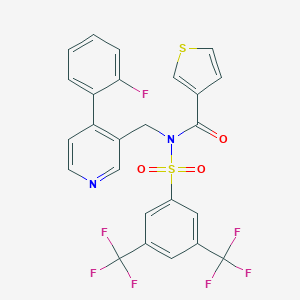 CHEMBL2435914 CHEMBL2435914 | C25H15F7N2O3S2 | 588.515 | 12 / 0 | 6.1 | No |
| 184154 | 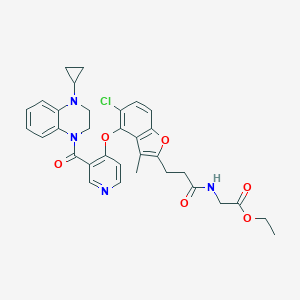 CHEMBL3290737 CHEMBL3290737 | C33H33ClN4O6 | 617.099 | 8 / 1 | 5.1 | No |
| 184690 | 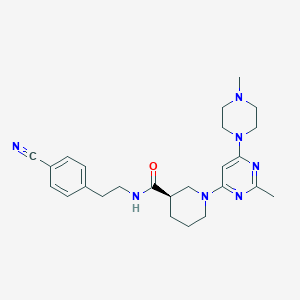 CHEMBL3234567 CHEMBL3234567 | C25H33N7O | 447.587 | 7 / 1 | 2.8 | Yes |
| 449120 | 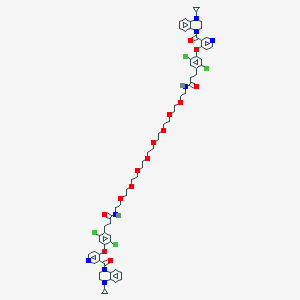 CHEMBL3421904 CHEMBL3421904 | C70H82Cl4N8O14 | 1401.27 | 18 / 2 | 7.8 | No |
| 189860 | 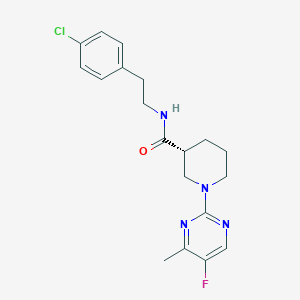 CHEMBL3234555 CHEMBL3234555 | C19H22ClFN4O | 376.86 | 5 / 1 | 3.5 | Yes |
| 563336 | 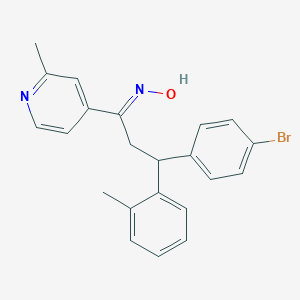 SCHEMBL10241033 SCHEMBL10241033 | C22H21BrN2O | 409.327 | 3 / 1 | 5.6 | No |
| 449722 | 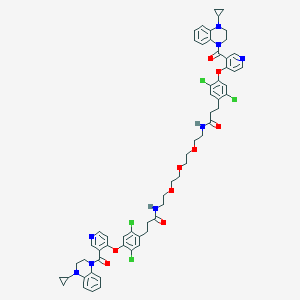 CHEMBL3421902 CHEMBL3421902 | C60H62Cl4N8O9 | 1181.0 | 13 / 2 | 8.6 | No |
| 563756 | 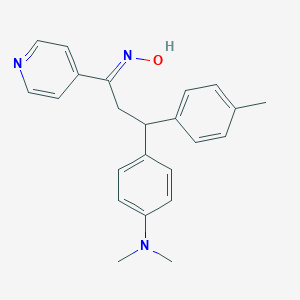 SCHEMBL364053 SCHEMBL364053 | C23H25N3O | 359.473 | 4 / 1 | 4.6 | Yes |
| 206456 | 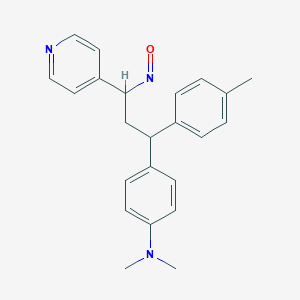 BDBM50437887 BDBM50437887 | C23H25N3O | 359.473 | 4 / 0 | 4.4 | Yes |
| 206932 | 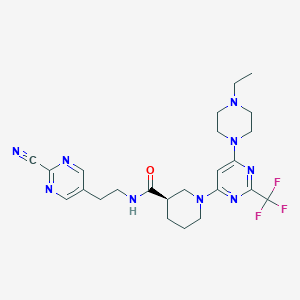 CHEMBL3234573 CHEMBL3234573 | C24H30F3N9O | 517.561 | 12 / 1 | 2.3 | No |
| 563832 | 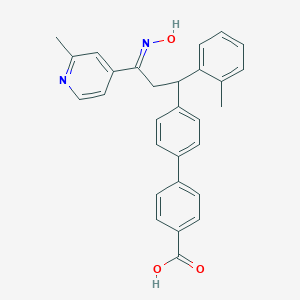 CHEMBL2407954 CHEMBL2407954 | C29H26N2O3 | 450.538 | 5 / 2 | 6.1 | No |
| 563833 |  SCHEMBL364196 SCHEMBL364196 | C29H26N2O3 | 450.538 | 5 / 2 | 6.1 | No |
| 563836 | 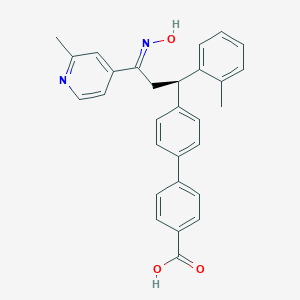 SCHEMBL362606 SCHEMBL362606 | C29H26N2O3 | 450.538 | 5 / 2 | 6.1 | No |
| 207566 |  CHEMBL3234571 CHEMBL3234571 | C24H29F3N8O | 502.546 | 11 / 1 | 2.6 | No |
| 449948 |  CHEMBL3421901 CHEMBL3421901 | C29H28Cl2N4O5 | 583.466 | 7 / 1 | 4.4 | No |
| 211992 | 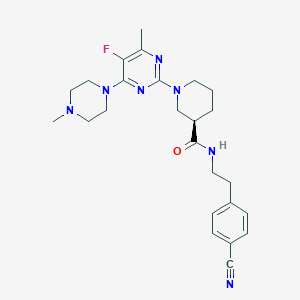 CHEMBL3234566 CHEMBL3234566 | C25H32FN7O | 465.577 | 8 / 1 | 2.9 | Yes |
| 213854 |  CHEMBL2181240 CHEMBL2181240 | C24H21Cl2N3O2 | 454.351 | 4 / 0 | 5.5 | No |
| 216593 | 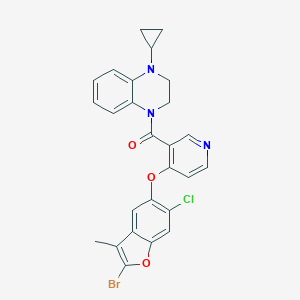 CHEMBL3290718 CHEMBL3290718 | C26H21BrClN3O3 | 538.826 | 5 / 0 | 6.3 | No |
| 217680 | 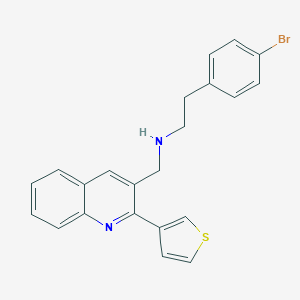 CHEMBL1257490 CHEMBL1257490 | C22H19BrN2S | 423.372 | 3 / 1 | 5.5 | No |
| 217808 | 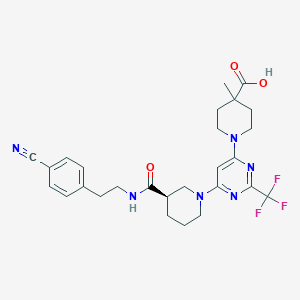 CHEMBL3234581 CHEMBL3234581 | C27H31F3N6O3 | 544.579 | 11 / 2 | 3.9 | No |
| 224025 | 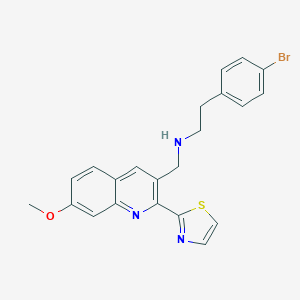 CHEMBL1258878 CHEMBL1258878 | C22H20BrN3OS | 454.386 | 5 / 1 | 4.9 | Yes |
| 224170 |  CHEMBL3234570 CHEMBL3234570 | C27H32F3N7O2 | 543.595 | 11 / 1 | 3.0 | No |
| 224614 |  BDBM50437877 BDBM50437877 | C28H31N3O3 | 457.574 | 6 / 1 | 4.9 | Yes |
| 224615 | 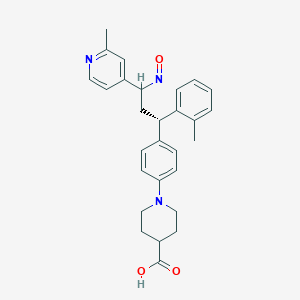 BDBM50437876 BDBM50437876 | C28H31N3O3 | 457.574 | 6 / 1 | 4.9 | Yes |
| 228768 |  INT-777 INT-777 | C27H46O5 | 450.66 | 5 / 4 | 4.9 | Yes |
zhanglab![]() zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218
zhanggroup.org | (734) 647-1549 | 100 Washtenaw Avenue, Ann Arbor, MI 48109-2218